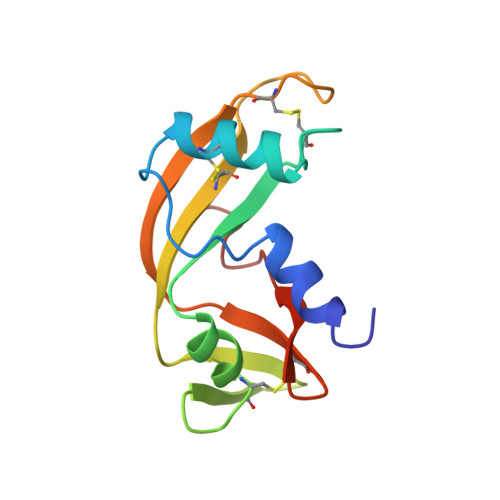
Three-dimensional solution structure of human angiogenin determined by 1H,15N-NMR spectroscopy--characterization of histidine protonation states and pKa values.
Lequin, O., Thuring, H., Robin, M., Lallemand, J.Y.(1997) Eur J Biochem 250: 712-726
- PubMed: 9461294 Search on PubMed
- DOI: https://doi.org/10.1111/j.1432-1033.1997.00712.x
- Primary Citation Related Structures:
1AWZ - PubMed Abstract:
Human angiogenin is a member of the pancreatic ribonuclease superfamily that induces blood vessel formation. Its three-dimensional solution structure has been determined to high resolution by heteronuclear NMR spectroscopy. 30 structures were calculated, based on a total of 1441 assigned NOE correlations, 64 coupling constants and 50 hydrogen bonds. The backbone atomic rms difference from the mean coordinates is 0.067 +/- 0.012 nm and 0.13 nm from the previously determined crystal structure. The side-chain of Gln117 was found to obstruct the active site as observed in the crystal state. There was no evidence of an alternative open form of angiogenin, although two sets of chemical shifts were observed for some residues, mainly around the active site and in the C-terminal segment. The topology of the ribonucleolytic active site is described with a particular emphasis on the conformation and protonation of active-site His residues. The side-chain of His114 adopts two main conformations in solution. In contrast to pancreatic ribonuclease A, His13 was shown to be more basic than His114, with pKa values of 6.65 and 6.05 respectively. The His47 residue is located in an environment very resistant to protonation with a pKa lower than 4.
- Département de Chimie-Synthèse Organique, URA 1308 du CNRS, Ecole Polytechnique, Palaiseau, France.
Organizational Affiliation: