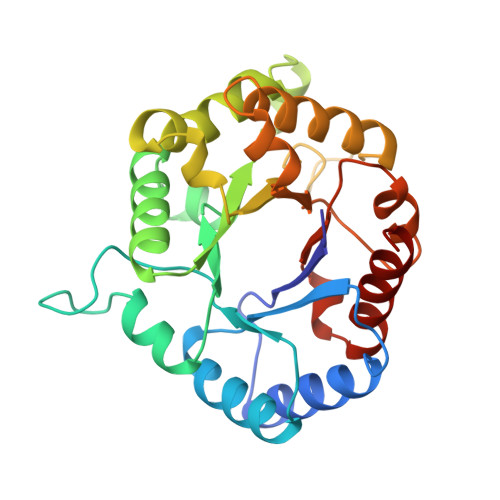
Triose-phosphate isomerase (TIM) of the psychrophilic bacterium Vibrio marinus. Kinetic and structural properties.
Alvarez, M., Zeelen, J.P., Mainfroid, V., Rentier-Delrue, F., Martial, J.A., Wyns, L., Wierenga, R.K., Maes, D.(1998) J Biological Chem 273: 2199-2206
- PubMed: 9442062 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.273.4.2199
- Primary Citation Related Structures:
1AW1, 1AW2 - PubMed Abstract:
The purification and characterization of triose-phosphate isomerase from the psychrophilic bacterium Vibrio marinus (vTIM) is described. Crystal structures of the vTIM-sulfate complex and the vTIM-2-phosphoglycolate complex (at a 2.7-A resolution) are also presented. The optimal growth temperature of Vibrio marinus is 15 degrees C. Stability studies show that vTIM is an unstable protein with a half-life of only 10 min at 25 degrees C. The vTIM sequence is most closely related to the sequence of Escherichia coli TIM (eTIM) (66% identity), and several unique structural features described for eTIM are also seen in vTIM, but eTIM is considerably more stable. The Td values of vTIM and eTIM, determined by calorimetric studies, are 41 and 54 degrees C, respectively. Amino acid sequence comparison reveals that vTIM has an alanine in loop 8 (at position 238), whereas all other TIM sequences known to date have a serine. The vTIM mutant A238S was produced and characterized. Compared with wild type, the catalytic efficiency of the A238S mutant is somewhat reduced, and its stability is considerably increased.
- Laboratoire de Biologie Moléculaire et de Génie Génétique, Université de Liège, Sart Tilman, Belgium.
Organizational Affiliation: