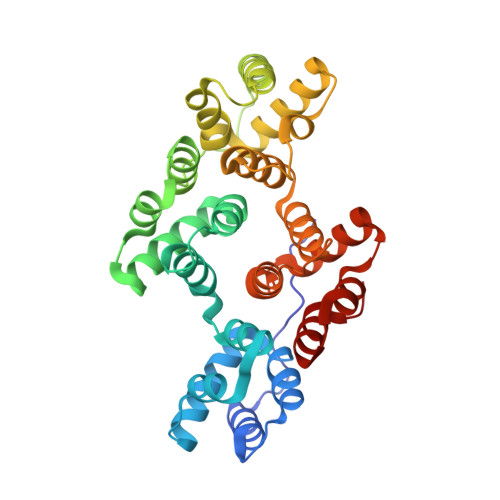
Crystal and molecular structure of human annexin V after refinement. Implications for structure, membrane binding and ion channel formation of the annexin family of proteins.
Huber, R., Berendes, R., Burger, A., Schneider, M., Karshikov, A., Luecke, H., Romisch, J., Paques, E.(1992) J Mol Biology 223: 683-704
- PubMed: 1311770
- DOI: https://doi.org/10.1016/0022-2836(92)90984-r
- Primary Citation of Related Structures:
1AVH, 1AVR, 1SAV - PubMed Abstract:
Two crystal forms (P6(3) and R3) of human annexin V have been crystallographically refined at 2.3 A and 2.0 A resolution to R-values of 0.184 and 0.174, respectively, applying very tight stereochemical restraints with deviations from ideal geometry of 0.01 A and 2 degrees. The three independent molecules (2 in P6(3), 1 in R3) are similar, with deviations in C alpha positions of 0.6 A. The polypeptide chain of 320 amino acid residues is folded into a planar cyclic arrangement of four repeats. The repeats have similar structures of five alpha-helical segments wound into a right-handed compact superhelix. Three calcium ion sites in repeats I, II and IV and two lanthanum ion sites in repeat I have been found in the R3 crystals. They are located at the convex face of the molecule opposite the N terminus. Repeat III has a different conformation at this site and no calcium bound. The calcium sites are similar to the phospholipase A2 calcium-binding site, suggesting analogy also in phospholipid interaction. The center of the molecule is formed by a channel of polar charged residues, which also harbors a chain of ordered water molecules conserved in the different crystal forms. Comparison with amino acid sequences of other annexins shows a high degree of similarity between them. Long insertions are found only at the N termini. Most conserved are the residues forming the metal-binding sites and the polar channel. Annexins V and VII form voltage-gated calcium ion channels when bound to membranes in vitro. We suggest that annexins bind with their convex face to membranes, causing local disorder and permeability of the phospholipid bilayers. Annexins are Janus-faced proteins that face phospholipid and water and mediate calcium transport.
- Max-Planck-Institut für Biochemie, Martinsried, Germany.
Organizational Affiliation: