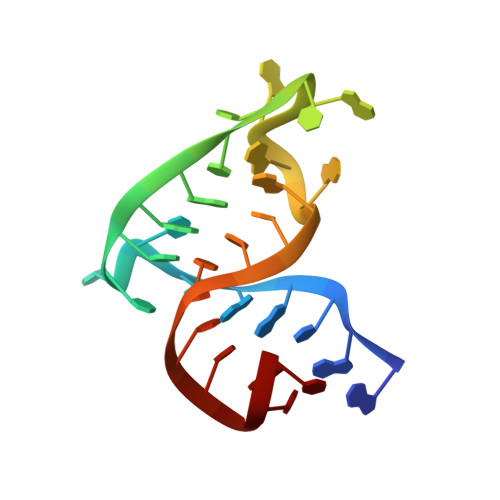
The structure of the human immunodeficiency virus type-1 TAR RNA reveals principles of RNA recognition by Tat protein.
Aboul-ela, F., Karn, J., Varani, G.(1995) J Mol Biology 253: 313-332
- PubMed: 7563092 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1995.0555
- Primary Citation Related Structures:
1ARJ - PubMed Abstract:
The human immunodeficiency virus type-1 (HIV-1) Tat protein stimulates transcriptional elongation. Tat is introduced to the transcription machinery by binding to the transactivation response region (TAR) RNA stem-loop encoded by the 5' leader sequence found on all HIV-1 mRNAs. We have used multidimensional heteronuclear NMR to determine the structure of the TAR RNA in the presence of the ADP-1 polypeptide, a 37-mer that carries the minimal RNA recognition region of the Tat protein and closely mimics Tat binding specificity. In the presence of a variety of ligands, including ADP-1, related basic peptides and the amino acid derivative argininamide, the bulge region of TAR undergoes a local conformational rearrangement and forms a more stable structure. The structure of TAR in the bound form has been determined from over 1000 NMR-derived constraints. The U23 residue at the 5' end of the bulge is positioned near G26 and A27 in the major groove, rather than stacked on A22 as in the free TAR. U23 and G26 are brought into close proximity by contacts to the guanidinium group and side-chain amide group of a common arginine residue. However, the interaction of this guanidinium group with TAR is not the only source of binding specificity. Besides NOEs to the arginine residue participating in the conformational change, ADP-1 shows additional intermolecular NOEs to TAR, suggesting that there are multiple points of contacts between TAR RNA and residues from the basic and core regions of Tat. These structural results provide important clues towards the identification of small molecular mass and/or peptidomimetic inhibitors of the essential Tat-TAR interaction.
- MRC Laboratory of Molecular Biology, Cambridge, UK.
Organizational Affiliation: