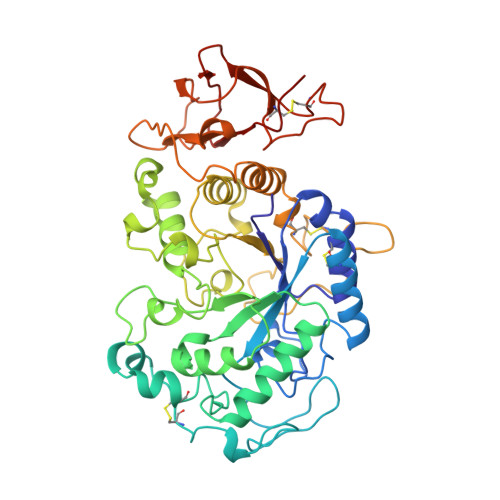
Crystal structures of the psychrophilic alpha-amylase from Alteromonas haloplanctis in its native form and complexed with an inhibitor.
Aghajari, N., Feller, G., Gerday, C., Haser, R.(1998) Protein Sci 7: 564-572
- PubMed: 9541387
- DOI: https://doi.org/10.1002/pro.5560070304
- Primary Citation of Related Structures:
1AQH, 1AQM - PubMed Abstract:
Alteromonas haloplanctis is a bacterium that flourishes in Antarctic sea-water and it is considered as an extreme psychrophile. We have determined the crystal structures of the alpha-amylase (AHA) secreted by this bacterium, in its native state to 2.0 angstroms resolution as well as in complex with Tris to 1.85 angstroms resolution. The structure of AHA, which is the first experimentally determined three-dimensional structure of a psychrophilic enzyme, resembles those of other known alpha-amylases of various origins with a surprisingly greatest similarity to mammalian alpha-amylases. AHA contains a chloride ion which activates the hydrolytic cleavage of substrate alpha-1,4-glycosidic bonds. The chloride binding site is situated approximately 5 angstroms from the active site which is characterized by a triad of acid residues (Asp 174, Glu 200, Asp 264). These are all involved in firm binding of the Tris moiety. A reaction mechanism for substrate hydrolysis is proposed on the basis of the Tris inhibitor binding and the chloride activation. A trio of residues (Ser 303, His 337, Glu 19) having a striking spatial resemblance with serine-protease like catalytic triads was found approximately 22 angstroms from the active site. We found that this triad is equally present in other chloride dependent alpha-amylases, and suggest that it could be responsible for autoproteolytic events observed in solution for this cold adapted alpha-amylase.
- Laboratoire d'Architecture et Fonction des Macromolécules Biologiques, UPR9039, Institut de Biologie Structurale et Microbiologie, IFR1, CNRS, Marseille, France.
Organizational Affiliation: