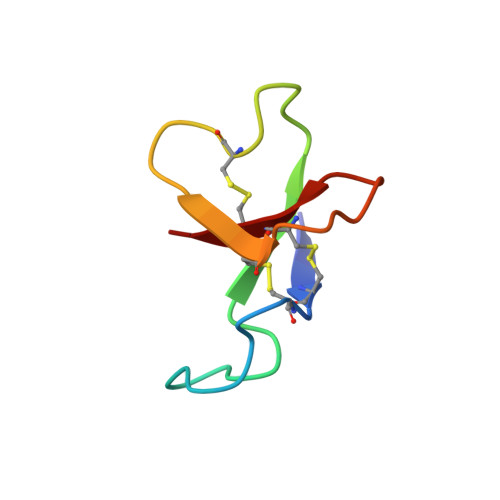
Solution structure of the cardiostimulant polypeptide anthopleurin-B and comparison with anthopleurin-A.
Monks, S.A., Pallaghy, P.K., Scanlon, M.J., Norton, R.S.(1995) Structure 3: 791-803
- PubMed: 7582896 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00214-3
- Primary Citation Related Structures:
1APF - PubMed Abstract:
The polypeptide anthopleurin-B (AP-B) is one of a number of related toxins produced by sea anemones. AP-B delays inactivation of the voltage-gated sodium channel of excitable tissue. In the mammalian heart, this effect is manifest as an increase in the force of contraction. As a result, there is interest in exploiting the anthopleurins as lead compounds in the design of novel cardiac stimulants. Essential to this endeavour is a high-resolution solution structure of the molecule describing the positions of functionally important side chains. AP-B exists in multiple conformations in solution as a result of cis-trans isomerization about the Gly40-Pro41 peptide bond. The solution structure of the major conformer of AP-B has been determined by two-dimensional 1H NMR at pH 4.5 and 25 degrees C. The core structure is a four-stranded, antiparallel beta-sheet (residues 2-4, 20-23, 34-37 and 45-48) and includes several beta-turns (6-9, 25-28, 30-33). Three loops connect the beta-strands, the longest and least well defined being the first loop, extending from residues 8-17. These features are shared by other members of this family of sea anemone toxins. The locations of a number of side chains which are important for the cardiac stimulatory activity of AP-B are well defined in the structures. We have described the solution structure of AP-B and compared it with that of AP-A, from which it differs by substitutions at seven amino acid positions. It shares an essentially identical fold with AP-A yet is about 10-fold more active. Comparison of the structures, particularly in the region of residues essential for activity, gives a clearer indication of the location and extent of the cardioactive pharmacophore in these polypeptides.
- NMR Laboratory, Biomolecular Research Institute, Parkville, Australia.
Organizational Affiliation: