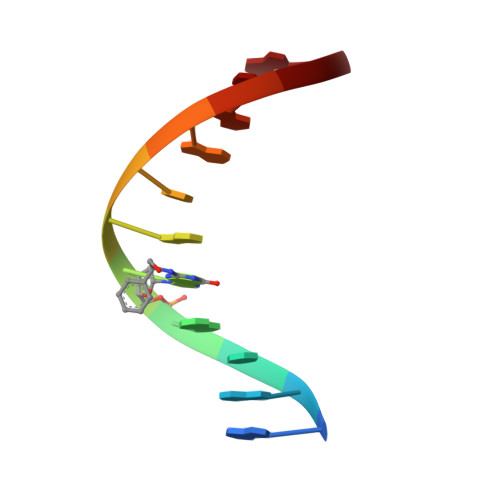
Styrene oxide adducts in an oligodeoxynucleotide containing the human N-ras codon 12: minor groove structures of the R(12,1)- and S(12,1)-alpha-(N2-guanyl) stereoisomers determined by 1H nuclear magnetic resonance.
Setayesh, F.R., DeCorte, B.L., Horton, P., Harris, C.M., Harris, T.M., Stone, M.P.(1998) Chem Res Toxicol 11: 766-777
- PubMed: 9671539
- DOI: https://doi.org/10.1021/tx9800147
- Primary Citation Related Structures:
1AP1 - PubMed Abstract:
The R- and S-alpha-(N2-guanyl)styrene oxide (SO) adducts at X5 in d(G1G2C3A4X5G6T7G8G9T10G11).d(C12A13C14C15A16C17C18T19G20C21C22 ), encompassing codon 12 of the human N-ras protooncogene (underlined), were examined using 1H NMR spectroscopy. These were the R(12,1) and S(12,1) adducts, indicating the location of the R or S adduct at the first position of codon 12. These differed from the R- and S(12, 2)-alpha-SO adducts [Zegar, I. S., Setayesh, F. R., DeCorte, B. L., Harris, C. M., Harris, T. M., and Stone, M. P. (1996) Biochemistry 35, 4334-4348] in that the base pair 5' to the lesion was changed from G.C to A.T, while the base pair 3' to the lesion was changed from T.A to G.C. Comparison of the R- and S(12,1) adducts with the R- and S(12,2) adducts allowed the effects of flanking bases on the conformations of the alpha-SO adducts to be examined. This change in flanking base affected the R-SO lesion. The R(12,1) adduct structure was disordered at the adduct site, and a refined structure could not be obtained. NOE and chemical shift data suggested that the styrenyl moiety was oriented in the minor groove and in the 3'-direction from the site of adduction. In contrast, this change in flanking base did not affect the S-SO lesion. The S(12,1) adduct yielded a refined structure, with the styrenyl moiety edgewise in the minor groove and oriented in the 5'-direction relative to the site of adduction. A total of 232 interproton distances, including 13 styrene-DNA distances, were obtained. A total of 12 NOE-restrained molecular dynamics calculations converged with pairwise root-mean-square deviation of 1.10 A. The sixth-root residual index between calculated and experimental NOE intensities was 8.0 x 10(-)2 A. The styrene aromatic protons appeared as three resonances, suggesting rapid rotation. The possibility of a hydrogen bond between the styrene hydroxyl and C18 O2 in the S(12,1) adduct could not be confirmed. This work illustrates the dual roles of stereochemistry and sequence in modulating the properties of guanine N2 alpha-SO adducts.
- Department of Chemistry and Center in Molecular Toxicology, Vanderbilt University, Nashville, Tennessee 37235, USA.
Organizational Affiliation: