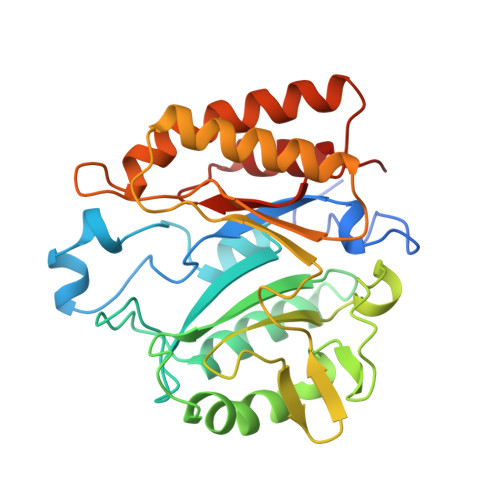
Crystal structure of the phosphatidylinositol-specific phospholipase C from the human pathogen Listeria monocytogenes.
Moser, J., Gerstel, B., Meyer, J.E., Chakraborty, T., Wehland, J., Heinz, D.W.(1997) J Mol Biology 273: 269-282
- PubMed: 9367761 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1290
- Primary Citation Related Structures:
1AOD, 2PLC - PubMed Abstract:
The X-ray crystal structure of the phosphatidylinositol-specific phospholipase C (PI-PLC) from the human pathogen Listeria monocytogenes has been determined both in free form at 2.0 A resolution, and in complex with the competitive inhibitor myo-inositol at 2.6 A resolution. The structure was solved by a combination of molecular replacement using the structure of Bacillus cereus PI-PLC and single isomorphous replacement. The enzyme consists of a single (beta alpha)8-barrel domain with the active site located at the C-terminal side of the beta-barrel. Unlike other (beta alpha)8-barrels, the barrel in PI-PLC is open because it lacks hydrogen bonding interactions between beta-strands V and VI. myo-Inositol binds to the active site pocket by making specific hydrogen bonding interactions with a number of charged amino acid side-chains as well as a coplanar stacking interaction with a tyrosine residue. Despite a relatively low sequence identity of approximately 24%, the structure is highly homologous to that of B.cereus PI-PLC with an r.m.s. deviation for 228 common C alpha positions of 1.46 A. Larger differences are found for loop regions that accommodate most of the numerous amino acid insertions and deletions. The active site pocket is also well conserved with only two amino acid replacements directly implicated in inositol binding.
- Institut für Organische Chemie und Biochemie, Universität Freiburg, Germany.
Organizational Affiliation: