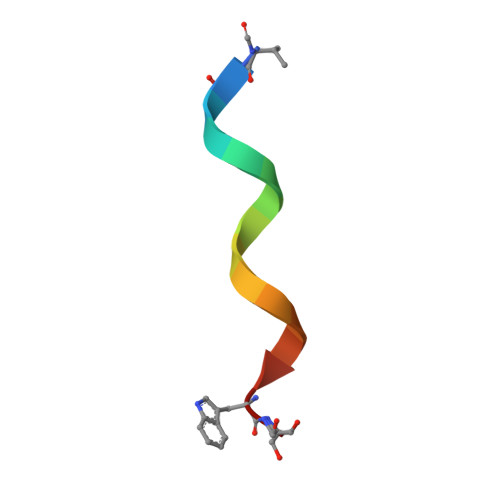
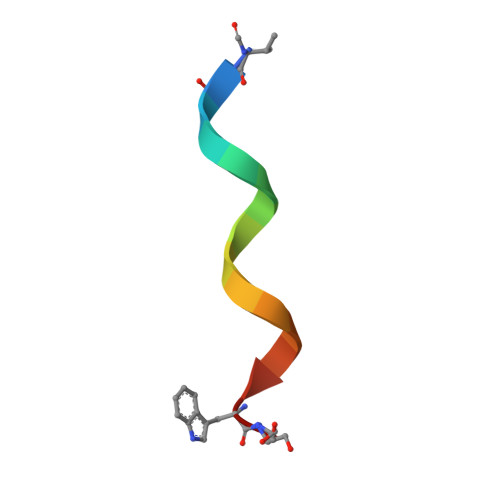
Heterodimer Formation and Crystal Nucleation of Gramicidin D
Burkhart, B.M., Gassman, R.M., Langs, D.A., Pangborn, W.A., Duax, W.L.(1998) Biophys J 75: 2135
- PubMed: 9788907 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/S0006-3495(98)77656-8
- Primary Citation Related Structures:
1AL4, 1ALX, 1ALZ - PubMed Abstract:
The linear pentadecapeptide antibiotic gramicidin D is a heterogeneous mixture of six components. Precise refinements of three-dimensional structures of naturally occurring gramicidin D in crystals obtained from methanol, ethanol, and n-propanol demonstrate the unexpected presence of stable left-handed antiparallel double-helical heterodimers that vary with the crystallization solvent. The side chains of Trp residues in the three structures exhibit sequence-specific patterns of conformational preference. Tyr substitution for Trp at position 11 appears to favor beta ribbon formation and stabilization of the antiparallel double helix that acts as a template for gramicidin folding and nucleation of different crystal forms. The fact that a minor component in a heterogeneous mixture influences aggregation and crystal nucleation has potential applications to other systems in which anomalous behavior is exhibited by aggregation of apparently homogeneous materials, such as the enigmatic behavior of prion proteins.
- Hauptman-Woodward Medical Research Institute, Inc., Buffalo, New York 14203-1196, USA.
Organizational Affiliation: