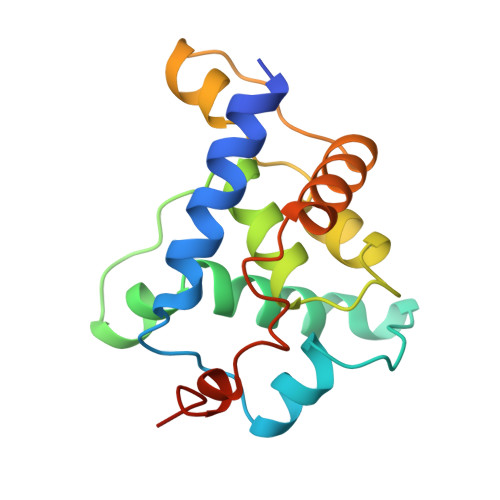
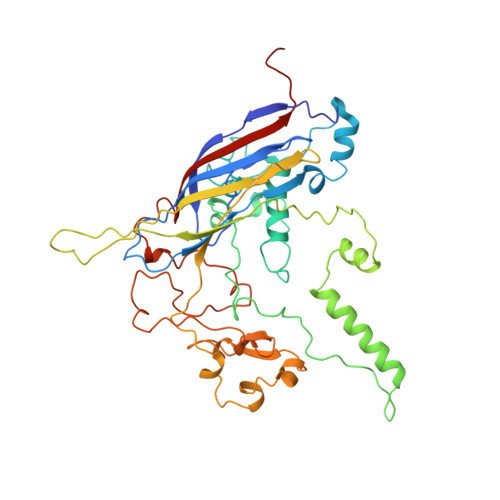
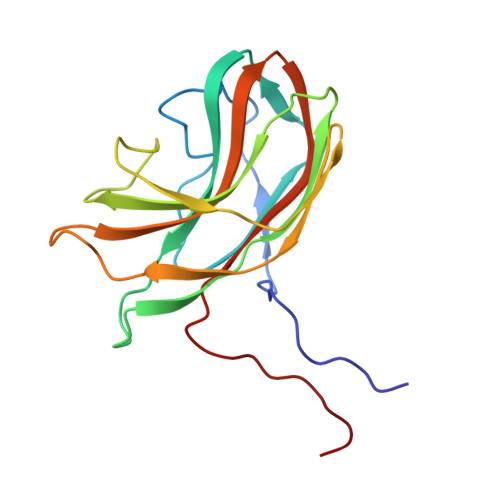
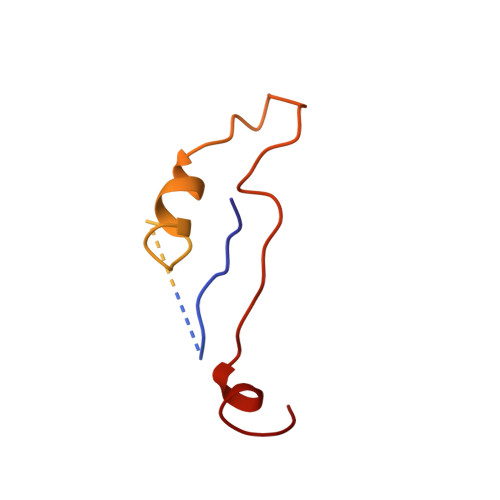
Structure of a viral procapsid with molecular scaffolding.
Dokland, T., McKenna, R., Ilag, L.L., Bowman, B.R., Incardona, N.L., Fane, B.A., Rossmann, M.G.(1997) Nature 389: 308-313
- PubMed: 9305849 Search on PubMed
- DOI: https://doi.org/10.1038/38537
- Primary Citation Related Structures:
1AL0 - PubMed Abstract:
The assembly of a macromolecular structure proceeds along an ordered morphogenetic pathway, and is accomplished by the switching of proteins between discrete conformations as they are added to the nascent assembly. Scaffolding proteins often play a catalytic role in the assembly process, rather like molecular chaperones. Although macromolecular assembly processes are fundamental to all biological systems, they have been characterized most thoroughly in viral systems, such as the icosahedral Escherichia coli bacteriophage phiX174. The phiX174 virion contains the proteins F, G, H and J. During assembly, two scaffoldingproteins B and D are required for the formation of a 108S, 360-A-diameter procapsid from pentameric precursors containing the F, G and H proteins. The procapsid contains 240 copies of protein D, forming an external scaffold, and 60 copies each of the internal scaffolding protein B, the capsid protein F, and the spike protein G. Maturation involves packaging of DNA and J proteins and loss of protein B, producing a 132S intermediate. Subsequent removal of the external scaffold yields the mature virion. Both the F and G proteins have the eight-stranded antiparallel beta-sandwich motif common to many plant and animal viruses. Here we describe the structure of a procapsid-like particle at 3.5-A resolution, showing how the scaffolding proteins coordinate assembly of the virus by interactions with the F and G proteins, and showing that the F protein undergoes conformational changes during capsid maturation.
- Department of Biological Sciences, Purdue University, West Lafayette, Indiana 47907-1392, USA.
Organizational Affiliation: