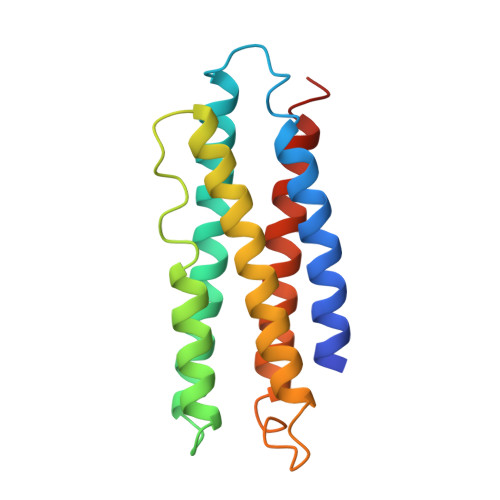
Molecular structure of an apolipoprotein determined at 2.5-A resolution.
Breiter, D.R., Kanost, M.R., Benning, M.M., Wesenberg, G., Law, J.H., Wells, M.A., Rayment, I., Holden, H.M.(1991) Biochemistry 30: 603-608
- PubMed: 1988048 Search on PubMed
- DOI: https://doi.org/10.1021/bi00217a002
- Primary Citation Related Structures:
1AEP - PubMed Abstract:
The three-dimensional structure of an apolipoprotein isolated from the African migratory locust Locusta migratoria has been determined by X-ray analysis to a resolution of 2.5 A. The overall molecular architecture of this protein consists of five long alpha-helices connected by short loops. As predicted from amino acid sequence analyses, these helices are distinctly amphiphilic with the hydrophobic residues pointing in toward the interior of the protein and the hydrophilic side chains facing outward. The molecule falls into the general category of up-and-down alpha-helical bundles as previously observed, for example, in cytochrome c'. Although the structure shows the presence of five long amphiphilic alpha-helices, the alpha-helical moment and hydrophobicity of the entire molecule fall into the range found for normal globular proteins. Thus, in order for the amphiphilic helices to play a role in the binding of the protein to a lipid surface, there must be a structural reorganization of the protein which exposes the hydrophobic interior to the lipid surface. The three-dimensional motif of this apolipoprotein is compatible with a model in which the molecule binds to the lipid surface via a relatively nonpolar end and then spreads on the surface in such a way as to cause the hydrophobic side chains of the helices to come in contact with the lipid surface, the charged and polar residues to remain in contact with water, and the overall helical motif of the protein to be maintained.
- Department of Chemistry, University of Wisconsin, Madison 53705.
Organizational Affiliation: