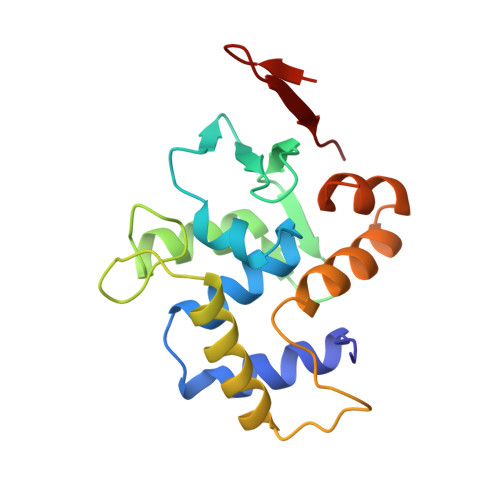
Flexibility in DNA recombination: structure of the lambda integrase catalytic core.
Kwon, H.J., Tirumalai, R., Landy, A., Ellenberger, T.(1997) Science 276: 126-131
- PubMed: 9082984 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.276.5309.126
- Primary Citation Related Structures:
1AE9 - PubMed Abstract:
Lambda integrase is archetypic of site-specific recombinases that catalyze intermolecular DNA rearrangements without energetic input. DNA cleavage, strand exchange, and religation steps are linked by a covalent phosphotyrosine intermediate in which Tyr342 is attached to the 3'-phosphate of the DNA cut site. The 1.9 angstrom crystal structure of the integrase catalytic domain reveals a protein fold that is conserved in organisms ranging from archaebacteria to yeast and that suggests a model for interaction with target DNA. The attacking Tyr342 nucleophile is located on a flexible loop about 20 angstroms from a basic groove that contains all the other catalytically essential residues. This bipartite active site can account for several apparently paradoxical features of integrase family recombinases, including the capacity for both cis and trans cleavage of DNA.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston MA 02115, USA.
Organizational Affiliation: