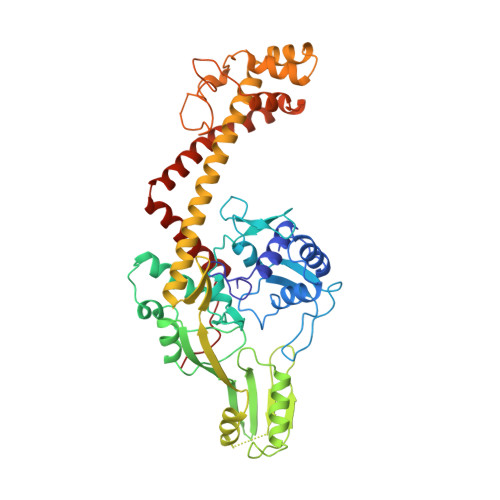
Crystal structure of the breakage-reunion domain of DNA gyrase.
Cabral, J.H., Jackson, A.P., Smith, C.V., Shikotra, N., Maxwell, A., Liddington, R.C.(1997) Nature 388: 903-906
- PubMed: 9278055 Search on PubMed
- DOI: https://doi.org/10.1038/42294
- Primary Citation Related Structures:
1AB4 - PubMed Abstract:
DNA gyrase is a type II DNA topoisomerase from bacteria that introduces supercoils into DNA. It catalyses the breakage of a DNA duplex (the G segment), the passage of another segment (the T segment) through the break, and then the reunification of the break. This activity involves the opening and dosing of a series of molecular 'gates' which is coupled to ATP hydrolysis. Here we present the crystal structure of the 'breakage-reunion' domain of the gyrase at 2.8 A resolution. Comparison of the structure of this 59K (relative molecular mass, 59,000) domain with that of a 92K fragment of yeast topoisomerase II reveals a very different quaternary organization, and we propose that the two structures represent two principal conformations that participate in the enzymatic pathway. The gyrase structure reveals a new dimer contact with a grooved concave surface for binding the G segment and a cluster of conserved charged residues surrounding the active-site tyrosines. It also shows how breakage of the G segment can occur and, together with the topoisomerase II structure, suggests a pathway by which the T segment can be released through the second gate of the enzyme. Mutations that confer resistance to the quinolone antibacterial agents cluster at the new dimer interface, indicating how these drugs might interact with the gyrase-DNA complex.
- Department of Biochemistry, University of Leicester, UK.
Organizational Affiliation: