DNA binding and cleavage by the nuclear intron-encoded homing endonuclease I-PpoI.
Flick, K.E., Jurica, M.S., Monnat Jr., R.J., Stoddard, B.L.(1998) Nature 394: 96-101
- PubMed: 9665136 Search on PubMed
- DOI: https://doi.org/10.1038/27952
- Primary Citation Related Structures:
1A73, 1A74, 1IPP - PubMed Abstract:
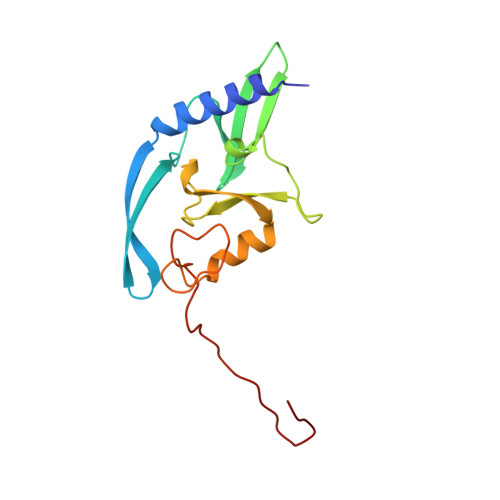
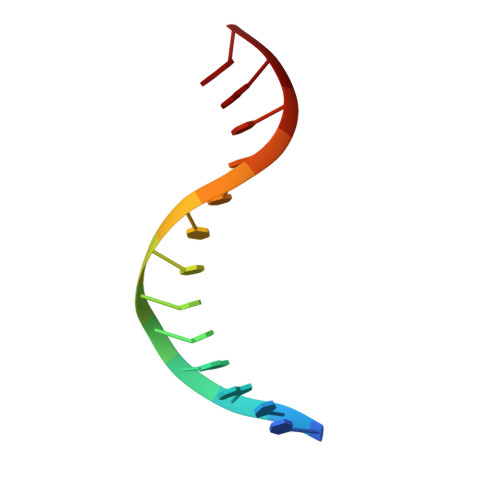
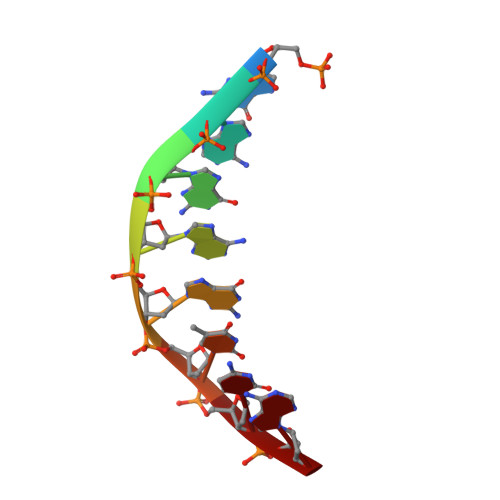
Homing endonucleases are a diverse collection of proteins that are encoded by genes with mobile, self-splicing introns. They have also been identified in self-splicing inteins (protein introns). These enzymes promote the movement of the DNA sequences that encode them from one chromosome location to another; they do this by making a site-specific double-strand break at a target site in an allele that lacks the corresponding mobile intron. The target sites recognized by these small endonucleases are generally long (14-44 base pairs). Four families of homing endonucleases have been identified, including the LAGLIDADG, the His-Cys box, the GIY-YIG and the H-N-H endonucleases. The first identified His-Cys box homing endonuclease was I-PpoI from the slime mould Physarum polycephalum. Its gene resides in one of only a few nuclear introns known to exhibit genetic mobility. Here we report the structure of the I-PpoI homing endonuclease bound to homing-site DNA determined to 1.8 A resolution. I-PpoI displays an elongated fold of dimensions 25 x 35 x 80 A, with mixed alpha/beta topology. Each I-PpoI monomer contains three antiparallel beta-sheets flanked by two long alpha-helices and a long carboxy-terminal tail, and is stabilized by two bound zinc ions 15 A apart. The enzyme possesses a new zinc-bound fold and endonuclease active site. The structure has been determined in both uncleaved substrate and cleaved product complexes.
- Fred Hutchinson Cancer Research Center, University of Washington, Seattle 98109, USA.
Organizational Affiliation: