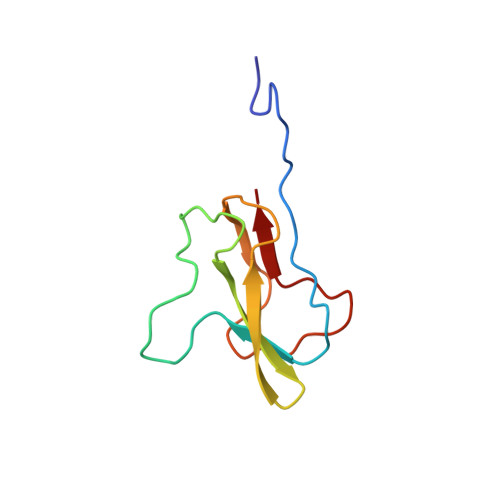
Structure of the carboxy-terminal fragment of the apo-biotin carboxyl carrier subunit of Escherichia coli acetyl-CoA carboxylase.
Yao, X., Wei, D., Soden Jr., C., Summers, M.F., Beckett, D.(1997) Biochemistry 36: 15089-15100
- PubMed: 9398236
- DOI: https://doi.org/10.1021/bi971485f
- Primary Citation Related Structures:
1A6X - PubMed Abstract:
The biotin carboxyl carrier protein (BCCP) is a subunit of acetyl-CoA carboxylase, a biotin-dependent enzyme that catalyzes the first committed step of fatty acid biosynthesis. In its functional cycle the biotin carboxyl carrier protein engages in heterologous protein-protein interactions with three distinct partners, depending on its state of posttranslational modification. Apo-BCCP interacts specifically with the biotin holoenzyme synthetase, BirA, which results in the posttranslational attachment of biotin to an essential lysine residue on BCCP. Holo-BCCP then interacts with the biotin carboxylase subunit, which leads to the addition of the carboxylate group of bicarbonate to biotin. Finally, the carboxybiotinylated form of BCCP interacts with transcarboxylase in the conversion of acetyl-CoA to malonyl-CoA. The determinants of protein-protein interaction specificity in this system are unknown. One hypothesis is that posttranslational modification of BCCP may result in conformational changes that regulate specific protein-protein interactions. To test this hypothesis, we have determined the NMR solution structure of the unbiotinylated form of an 87 residue C-terminal domain fragment of BCCP (apoBCCP87) from Escherichia coli acetyl-CoA carboxylase and compared this structure with the high-resolution structure of the biotinylated form that was recently solved by X-ray crystallographic techniques. Although the overall folding of the two proteins is highly similar, small structural differences are apparent for residues of the biotin-binding loop that may be important for mediating specific protein-protein interactions.
- Department of Chemistry and Biochemistry and Howard Hughes Medical Institute, University of Maryland Baltimore County, 1000 Hilltop Circle, Baltimore, Maryland 21250, USA.
Organizational Affiliation: