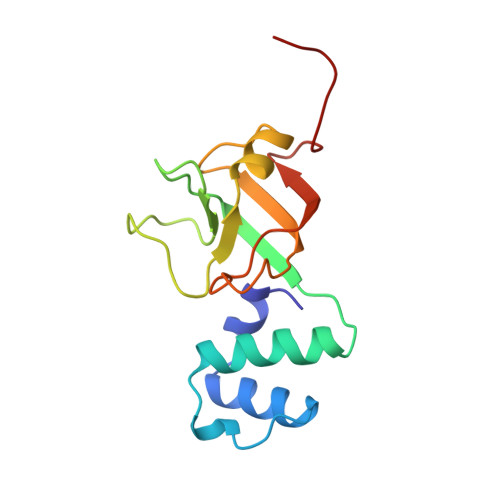
The NMR structure of the RNA binding domain of E. coli rho factor suggests possible RNA-protein interactions.
Briercheck, D.M., Wood, T.C., Allison, T.J., Richardson, J.P., Rule, G.S.(1998) Nat Struct Biol 5: 393-399
- PubMed: 9587002 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0598-393
- Primary Citation Related Structures:
1A63 - PubMed Abstract:
Rho protein is an essential hexameric RNA-DNA helicase that binds nascent mRNA transcripts and terminates transcription in a wide variety of eubacterial species. The NMR solution structure of the RNA binding domain of rho, rho130, is presented. This structure consists of two sub-domains, an N-terminal three-helix bundle and a C-terminal beta-barrel that is structurally similar to the oligosaccharide/oligonucleotide binding (OB) fold. Chemical shift changes of rho130 upon RNA binding and previous mutagenetic analyses of intact rho suggest that residues Asp 60, Phe 62, Phe 64, and Arg 66 are critical for binding and support the hypothesis that ssRNA/ssDNA binding is localized in the beta-barrel sub-domain. On the basis of these studies and the tertiary structure of rho130, we propose that residues Asp 60, Phe 62, Phe 64, Arg 66, Tyr 80, Lys 105, and Arg 109 participate in RNA-protein interactions.
- Department of Biochemistry, University of Virginia School of Medicine, Charlottesville 22908, USA.
Organizational Affiliation: