Conservation of structure and mechanism between eukaryotic topoisomerase I and site-specific recombinases.
Cheng, C., Kussie, P., Pavletich, N., Shuman, S.(1998) Cell 92: 841-850
- PubMed: 9529259 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(00)81411-7
- Primary Citation Related Structures:
1A41 - PubMed Abstract:
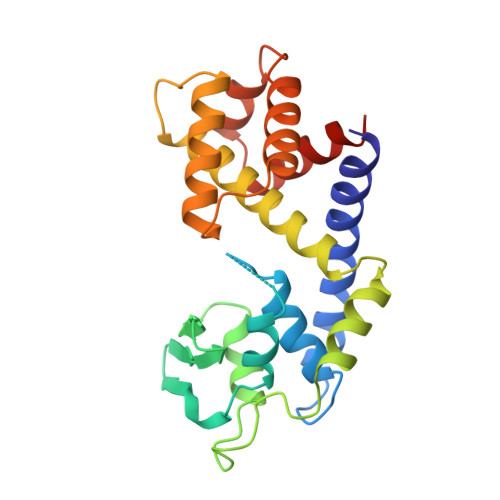
Vaccinia DNA topoisomerase breaks and rejoins DNA strands through a DNA-(3'-phosphotyrosyl)-enzyme intermediate. A C-terminal catalytic domain, Topo(81-314), suffices for transesterification chemistry. The domain contains a constellation of five amino acids, conserved in all eukaryotic type IB topoisomerases, that catalyzes attack of the tyrosine nucleophile on the scissile phosphate. The structure of the catalytic domain, consisting of ten alpha helices and a three-strand beta sheet, resembles the catalytic domains of site-specific recombinases that act via a topoisomerase IB-like mechanism. The topoisomerase catalytic pentad is conserved in the tertiary structures of the recombinases despite scant sequence similarity overall. This implies that the catalytic domains of type IB topoisomerases and recombinases derive from a common ancestral strand transferase.
- Molecular Biology Program, Sloan-Kettering Institute, New York, New York 10021, USA.
Organizational Affiliation: