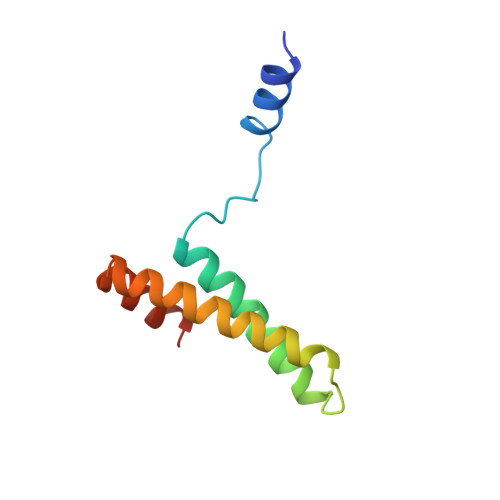
Conformational variability of the N-terminal helix in the structure of ribosomal protein S15.
Clemons Jr., W.M., Davies, C., White, S.W., Ramakrishnan, V.(1998) Structure 6: 429-438
- PubMed: 9562554 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(98)00045-8
- Primary Citation Related Structures:
1A32 - PubMed Abstract:
Ribosomal protein S15 is a primary RNA-binding protein that binds to the central domain of 16S rRNA. S15 also regulates its own synthesis by binding to its own mRNA. The binding sites for S15 on both mRNA and rRNA have been narrowed down to less than a hundred nucleotides each, making the protein an attractive candidate for the study of protein-RNA interactions. The crystal structure of S15 from Bacillus stearothermophilus has been solved to 2.1 A resolution. The structure consists of four alpha helices. Three of these helices form the core of the protein, while the N-terminal helix protrudes out from the body of the molecule to make contacts with a neighboring molecule in the crystal lattice. S15 contains a large conserved patch of basic residues which could provide a site for binding 16S rRNA. The conformation of the N-terminal alpha helix is quite different from that reported in a recent NMR structure of S15 from Thermus thermophilus. The intermolecular contacts that this alpha helix makes with a neighboring molecule in the crystal, however, closely resemble the intramolecular contacts that occur in the NMR structure. This conformational variability of the N-terminal helix has implications for the range of possible S15-RNA interactions. A large, conserved basic patch at one end of S15 and a cluster of conserved but exposed aromatic residues at the other end provide two possible RNA-binding sites on S15.
- Department of Biochemistry, University of Utah School of Medicine, Salt Lake City, UT 84132, USA.
Organizational Affiliation: