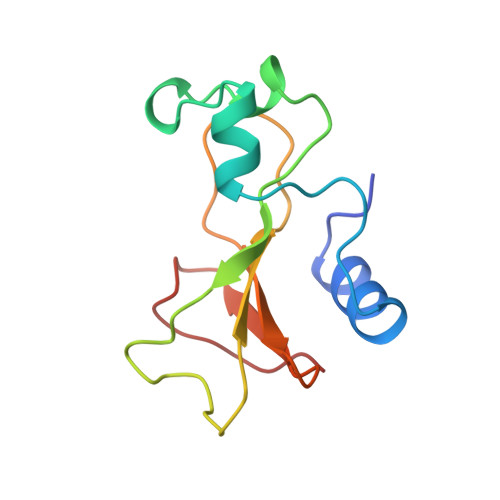
Refinement and structural analysis of barnase at 1.5 A resolution.
Martin, C., Richard, V., Salem, M., Hartley, R., Mauguen, Y.(1999) Acta Crystallogr D Biol Crystallogr 55: 386-398
- PubMed: 10089345 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444998010865
- Primary Citation Related Structures:
1A2P - PubMed Abstract:
The structure of Bacillus amyloliquefaciens ribonuclease (barnase), an extracellular 110-residue enzyme initially solved at 2.0 A resolution, has been refined at 1.5 A using synchrotron radiation and an imaging-plate scanner. Refinement with anisotropic atomic displacement parameters resulted in increased accuracy of the structure. The final model has a crystallographic R factor of 11.5% and an Rfree of 17.4%. The three independent molecules in the asymmetric unit, referred to as A, B and C, allowed detailed analysis of this final model and meaningful comparison with structures of barnase complexed either with nucleotide inhibitors or with its natural intracellular inhibitor, barstar. The analysis of the overall solvent structure revealed a similar number of water molecules associated with each barnase molecule; among these were 16 equivalent buried solvent molecules, the locations of which are discussed in detail and classified on the basis of their structural role. The importance of the water molecules' contribution to the barnase-barstar interaction is also highlighted. The high accuracy of the present analysis revealed the presence of a Zn2+ ion mediating the contacts between pairs of symmetry-related A, B or C molecules; such an ion had previously only been identified for pairs of C molecules.
- Laboratoire de Physique, CNRS, ERS 582, Centre d'Etudes Pharmaceutiques, 92296 Châtenay-Malabry CEDEX, France.
Organizational Affiliation: