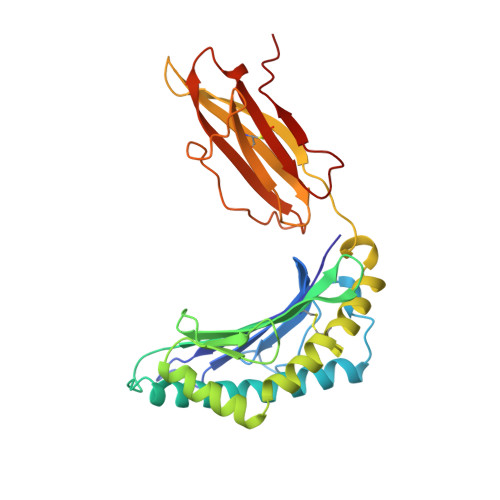
Bound water structure and polymorphic amino acids act together to allow the binding of different peptides to MHC class I HLA-B53.
Smith, K.J., Reid, S.W., Harlos, K., McMichael, A.J., Stuart, D.I., Bell, J.I., Jones, E.Y.(1996) Immunity 4: 215-228
- PubMed: 8624812 Search on PubMed
- DOI: https://doi.org/10.1016/s1074-7613(00)80430-6
- Primary Citation Related Structures:
1A1M, 1A1O - PubMed Abstract:
The structure of the human MHC class I molecule HLA-B53 complexed to two nonameric peptide epitopes (from the malaria parasite P. falciparum and the HIV2 gag protein) has been determined by X-ray crystallography at 2.3 angstrom resolution. The structures reveal the architecture of a Pro-specific B pocket common to many HLA-B alleles. Relative to other alleles, the B53 peptide-binding groove is widened by a significant (up to 1.25 angstrom) shift in the position of the alpha 1 helix. Within this groove, bound water molecules, acting in concert with the side chains of polymorphic residues, provide the functional malleability of the MHC, which enables the high affinity/low specificity binding of multiple peptide epitopes.
- Nuffield Department of Clinical Medicine, Institute of Molecular Medicine, John Radcliffe Hospital, Oxford, United Kingdom.
Organizational Affiliation: