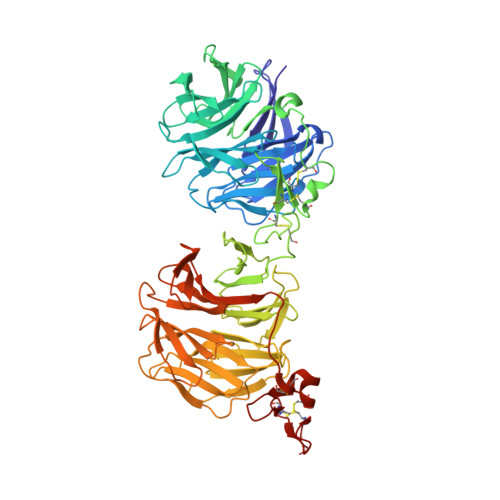
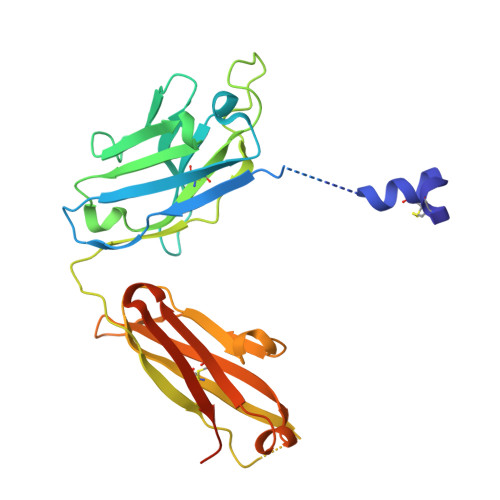
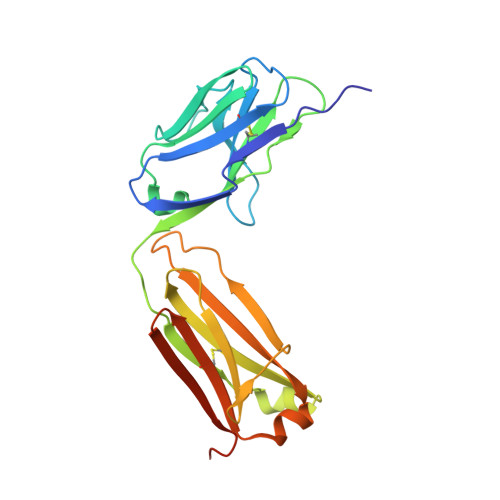
Structure of the Wnt-Frizzled-LRP6 initiation complex reveals the basis for coreceptor discrimination.
Tsutsumi, N., Hwang, S., Waghray, D., Hansen, S., Jude, K.M., Wang, N., Miao, Y., Glassman, C.R., Caveney, N.A., Janda, C.Y., Hannoush, R.N., Garcia, K.C.(2023) Proc Natl Acad Sci U S A 120: e2218238120-e2218238120
- PubMed: 36893265
- DOI: https://doi.org/10.1073/pnas.2218238120
- Primary Citation of Related Structures:
8CTG, 8FFE - PubMed Abstract:
Wnt morphogens are critical for embryonic development and tissue regeneration. Canonical Wnts form ternary receptor complexes composed of tissue-specific Frizzled (Fzd) receptors together with the shared LRP5/6 coreceptors to initiate β-catenin signaling. The cryo-EM structure of a ternary initiation complex of an affinity-matured XWnt8-Frizzled8-LRP6 complex elucidates the basis of coreceptor discrimination by canonical Wnts by means of their N termini and linker domains that engage the LRP6 E1E2 domain funnels. Chimeric Wnts bearing modular linker "grafts" were able to transfer LRP6 domain specificity between different Wnts and enable non-canonical Wnt5a to signal through the canonical pathway. Synthetic peptides comprising the linker domain serve as Wnt-specific antagonists. The structure of the ternary complex provides a topological blueprint for the orientation and proximity of Frizzled and LRP6 within the Wnt cell surface signalosome.
- HHMI, Stanford University School of Medicine, Stanford, CA 94305.
Organizational Affiliation: