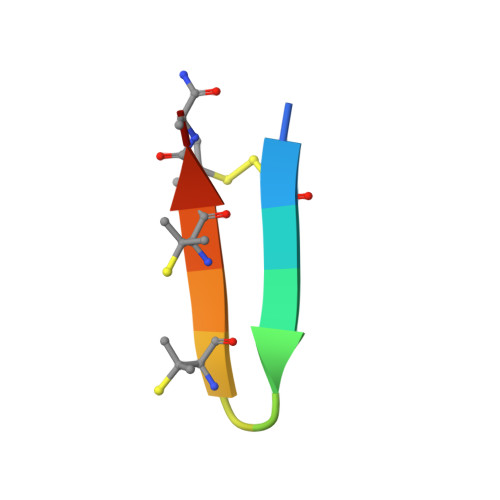
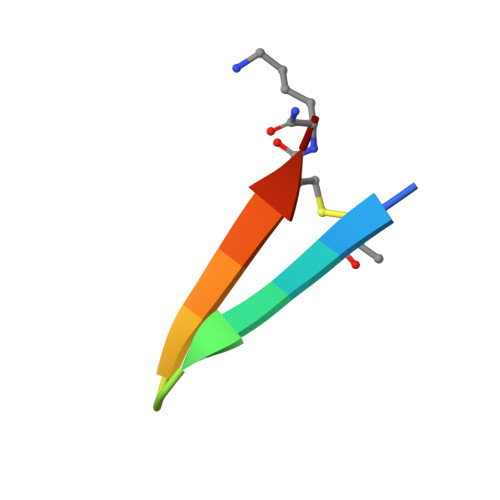
De novo design and directed folding of disulfide-bridged peptide heterodimers.
Yao, S., Moyer, A., Zheng, Y., Shen, Y., Meng, X., Yuan, C., Zhao, Y., Yao, H., Baker, D., Wu, C.(2022) Nat Commun 13: 1539-1539
- PubMed: 35318337
- DOI: https://doi.org/10.1038/s41467-022-29210-x
- Primary Citation of Related Structures:
7FB8, 7FBA - PubMed Abstract:
Peptide heterodimers are prevalent in nature, which are not only functional macromolecules but molecular tools for chemical and synthetic biology. Computational methods have also been developed to design heterodimers of advanced functions. However, these peptide heterodimers are usually formed through noncovalent interactions, which are prone to dissociate and subject to concentration-dependent nonspecific aggregation. Heterodimers crosslinked with interchain disulfide bonds are more stable, but it represents a formidable challenge for both the computational design of heterodimers and the manipulation of disulfide pairing for heterodimer synthesis and applications. Here, we report the design, synthesis and application of interchain disulfide-bridged peptide heterodimers with mutual orthogonality by combining computational de novo designs with a directed disulfide pairing strategy. These heterodimers can be used as not only scaffolds for generating functional molecules but chemical tools or building blocks for protein labeling and construction of crosslinking hybrids. This study thus opens the door for using this unexplored dimeric structure space for many biological applications.
- Department of Chemistry, College of Chemistry and Chemical Engineering, The MOE Key Laboratory of Spectrochemical Analysis and Instrumentation, State Key Laboratory of Physical Chemistry of Solid Surfaces, Xiamen University, Xiamen, 361005, P.R. China.
Organizational Affiliation: