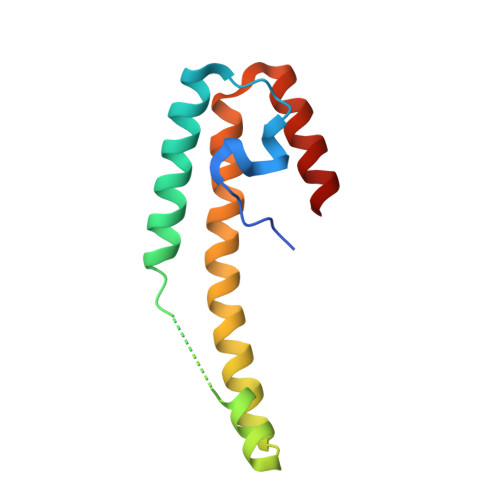
Structural investigation on SPI-6-associated Salmonella typhimurium VirG-like stress protein that promotes pathogen survival in macrophages.
Ray, S., Pandey, N.K., Kushwaha, G.S., Das, S., Ganguly, A.K., Vashi, N., Kumar, D., Suar, M., Bhavesh, N.S.(2022) Protein Sci 31: 835-849
- PubMed: 34997791
- DOI: https://doi.org/10.1002/pro.4272
- Primary Citation of Related Structures:
5IHF, 5IO8 - PubMed Abstract:
Enteric microbial pathogenesis, remarkably a complex process, is achieved by virulence factors encoded by genes located within regions of the bacterial genome termed pathogenicity islands. Salmonella pathogenicity islands (SPI) encodes proteins, that are essential virulence determinants for pathogen colonization and virulence. In addition to the well-characterized SPI-1 and SPI-2 proteins, which are required for bacterial invasion and intracellular replication, respectively, SPI-6 (formerly known as Salmonella enterica centisome 7 island [SCI]) encoding proteins are also known to play pivotal role in Salmonella pathogenesis. However, the underlying molecular mechanism of these proteins remained elusive. To gain molecular insights into SPI-6-associated proteins, in this study, a SPI-6 Salmonella typhimurium VirG-like protein (STV) is characterized using interdisciplinary experimental approaches including X-ray crystallography, nuclear magnetic resonance (NMR) spectroscopy and infection assays. The high-resolution crystal structure, determined by the single-wavelength anomalous dispersion (SAD) method, reveals that STV belongs to the LTxxQ motif family. Solution-state NMR spectroscopy studies reveal that STV form a dimer involving interconnected helices. Interestingly, functional studies show that STV influence pathogen persistence inside macrophages in vitro at later stages of infection. Altogether, our findings suggest that STV, a member of the LTxxQ stress protein family, modulates bacterial survival mechanism in macrophages through SPI-1 and SPI-2 genes, respectively.
- School of Biotechnology, Kalinga Institute of Industrial Technology (KIIT), Deemed to be University, Bhubaneswar, India.
Organizational Affiliation: