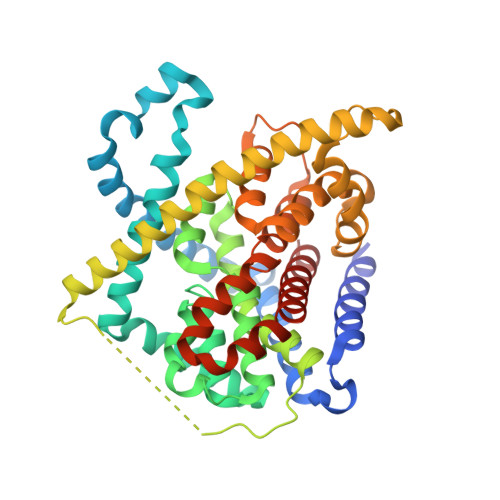
Crystal structure of a concentrative nucleoside transporter from Vibrio cholerae at 2.4A
Johnson, Z.L., Cheong, C.G., Lee, S.Y.(2012) Nature 483: 489-493
- PubMed: 22407322
- DOI: https://doi.org/10.1038/nature10882
- Primary Citation of Related Structures:
3TIJ - PubMed Abstract:
Nucleosides are required for DNA and RNA synthesis, and the nucleoside adenosine has a function in a variety of signalling processes. Transport of nucleosides across cell membranes provides the major source of nucleosides in many cell types and is also responsible for the termination of adenosine signalling. As a result of their hydrophilic nature, nucleosides require a specialized class of integral membrane proteins, known as nucleoside transporters (NTs), for specific transport across cell membranes. In addition to nucleosides, NTs are important determinants for the transport of nucleoside-derived drugs across cell membranes. A wide range of nucleoside-derived drugs, including anticancer drugs (such as Ara-C and gemcitabine) and antiviral drugs (such as zidovudine and ribavirin), have been shown to depend, at least in part, on NTs for transport across cell membranes. Concentrative nucleoside transporters, members of the solute carrier transporter superfamily SLC28, use an ion gradient in the active transport of both nucleosides and nucleoside-derived drugs against their chemical gradients. The structural basis for selective ion-coupled nucleoside transport by concentrative nucleoside transporters is unknown. Here we present the crystal structure of a concentrative nucleoside transporter from Vibrio cholerae in complex with uridine at 2.4 Å. Our functional data show that, like its human orthologues, the transporter uses a sodium-ion gradient for nucleoside transport. The structure reveals the overall architecture of this class of transporter, unravels the molecular determinants for nucleoside and sodium binding, and provides a framework for understanding the mechanism of nucleoside and nucleoside drug transport across cell membranes.
- Department of Biochemistry and Ion Channel Research Unit, Duke University Medical Center, 2 Genome Court, Durham, North Carolina 27710, USA.
Organizational Affiliation: