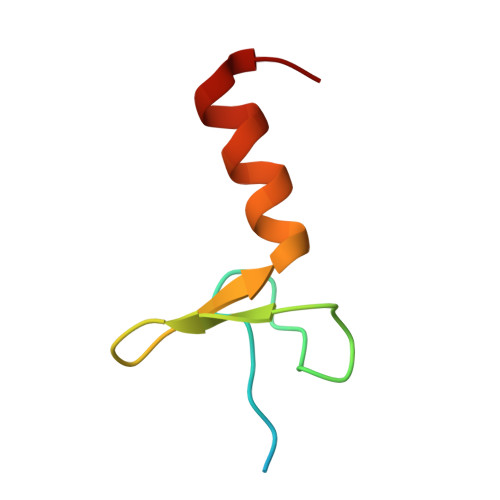
Structural Basis of SspB-tail Recognition by the Zinc Binding Domain of ClpX.
Park, E.Y., Lee, B.G., Hong, S.B., Kim, H.W., Jeon, H., Song, H.K.(2007) J Mol Biology 367: 514-526
- PubMed: 17258768
- DOI: https://doi.org/10.1016/j.jmb.2007.01.003
- Primary Citation of Related Structures:
2DS5, 2DS6, 2DS7, 2DS8 - PubMed Abstract:
The degradation of ssrA(AANDENYALAA)-tagged proteins in the bacterial cytosol is carried out by the ClpXP protease and is markedly stimulated by the SspB adaptor protein. It has previously been reported that the amino-terminal zinc-binding domain of ClpX (ZBD) is involved in complex formation with the SspB-tail (XB: ClpX-binding motif). In an effort to better understand the recognition of SspB by ClpX and the mechanism of delivery of ssrA-tagged substrates to ClpXP, we have determined the structures of ZBD alone at 1.5, 2.0, and 2.5 A resolution in each different crystal form and also in complex with XB peptide at 1.6 A resolution. The XB peptide forms an antiparallel beta-sheet with two beta-strands of ZBD, and the structure shows a 1:1 stoichiometric complex between ZBD and XB, suggesting that there are two independent SspB-tail-binding sites in ZBD. The high-resolution ZBD:XB complex structure, in combination with biochemical analyses, can account for key determinants in the recognition of the SspB-tail by ClpX and sheds light on the mechanism of delivery of target proteins to the prokaryotic degradation machine.
- School of Life Sciences and Biotechnology, Korea University, Seoul 136-701, Korea.
Organizational Affiliation: