SANS investigation of fungal loosenins reveals substrate-dependent impacts of protein action on the inter-microfibril arrangement of cellulosic substrates.
Dahiya, D., Peter-Szabo, Z., Senanayake, M., Pingali, S.V., Leite, W.C., Byrnes, J., Buchko, G.W., Sivan, P., Vilaplana, F., Master, E.R., O'Neill, H.(2025) Biotechnol Biofuels Bioprod 18: 27-27
- PubMed: 40022179
- DOI: https://doi.org/10.1186/s13068-025-02618-5
- Primary Citation of Related Structures:
9CE9 - PubMed Abstract:
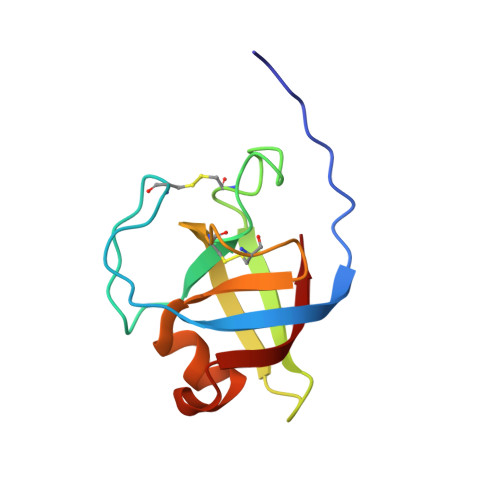
Microbial expansin-related proteins include fungal loosenins, which have been previously shown to disrupt cellulose networks and enhance the enzymatic conversion of cellulosic substrates. Despite showing beneficial impacts to cellulose processing, detailed characterization of cellulosic materials after loosenin treatment is lacking. In this study, small-angle neutron scattering (SANS) was used to investigate the effects of three recombinantly produced loosenins that originate from Phanerochaete carnosa, PcaLOOL7, PcaLOOL9, and PcaLOOL12, on the organization of holocellulose preparations from Eucalyptus and Spruce wood samples. Whereas the SANS analysis of Spruce holocellulose revealed an increase in inter-microfibril spacing of neighboring cellulose microfibrils following treatment with PcaLOOL12 and to a lesser extent PcaLOOL7, the analysis of Eucalyptus holocellulose revealed a reduction in the ordered arrangement of microfibrils following treatment with PcaLOOL12 and to a lesser extent PcaLOOL9. Parallel SEC-SAXS characterization of PcaLOOL7, PcaLOOL9, and PcaLOOL12 indicated the proteins likely function as monomers; moreover, all appear to retain a flexible disordered N-terminus and folded C-terminal region. The comparatively high impact of PcaLOOL12 motivated its NMR structural characterization, revealing a double-psi β-barrel (DPBB) domain surrounded by three α-helices-the largest nestled against the DPBB core and the other two part of loops extending from the core. The SANS analysis of PcaLOOL action on holocellulose samples confirms their ability to disrupt cellulose fiber networks and suggests a progression from reducing regular order in the microfibril arrangement to increasing inter-microfibril spacing. The most impactful PcaLOOL, PcaLOOL12, was previously observed to be the most highly expressed loosenin in P. carnosa. Its structural characterization herein reveals its stabilization through two disulfide linkages, and an extended N-terminal region distal to a negatively charged and surface accessible polysaccharide binding groove.
- Department of Bioproducts and Biosystems, Aalto University, Kemistintie 1, 02150, Espoo, Finland.
Organizational Affiliation: