Aphid BCR4 Structure and Activity Uncover a New Defensin Peptide Superfamily.
Loth, K., Parisot, N., Paquet, F., Terrasson, H., Sivignon, C., Rahioui, I., Ribeiro Lopes, M., Gaget, K., Duport, G., Delmas, A.F., Aucagne, V., Heddi, A., Calevro, F., da Silva, P.(2022) Int J Mol Sci 23
- PubMed: 36293341
- DOI: https://doi.org/10.3390/ijms232012480
- Primary Citation of Related Structures:
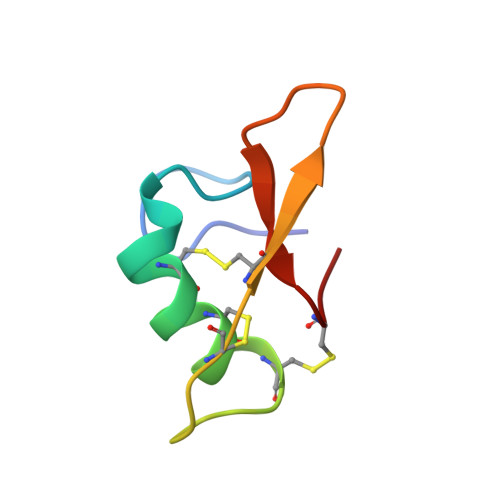
7PQW - PubMed Abstract:
Aphids (Hemiptera: Aphidoidea) are among the most detrimental insects for agricultural plants, and their management is a great challenge in agronomical research. A new class of proteins, called Bacteriocyte-specific Cysteine-Rich (BCR) peptides, provides an alternative to chemical insecticides for pest control. BCRs were initially identified in the pea aphid Acyrthosiphon pisum . They are small disulfide bond-rich proteins expressed exclusively in aphid bacteriocytes, the insect cells that host intracellular symbiotic bacteria. Here, we show that one of the A. pisum BCRs, BCR4, displays prominent insecticidal activity against the pea aphid, impairing insect survival and nymphal growth, providing evidence for its potential use as a new biopesticide. Our comparative genomics and phylogenetic analyses indicate that BCRs are restricted to the aphid lineage. The 3D structure of BCR4 reveals that this peptide belongs to an as-yet-unknown structural class of peptides and defines a new superfamily of defensins.
- Centre de Biophysique Moléculaire, CNRS UPR 4301, 45071 Orléans, France.
Organizational Affiliation: