Structural insight into DNA recognition by bacterial transcriptional regulators of the SorC/DeoR family.
Soltysova, M., Sieglova, I., Fabry, M., Brynda, J., Skerlova, J., Rezacova, P.(2021) Acta Crystallogr D Struct Biol 77: 1411-1424
- PubMed: 34726169
- DOI: https://doi.org/10.1107/S2059798321009633
- Primary Citation of Related Structures:
7BHY, 7OYK - PubMed Abstract:
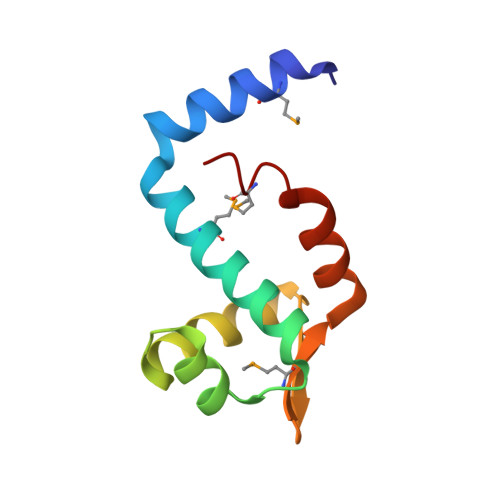
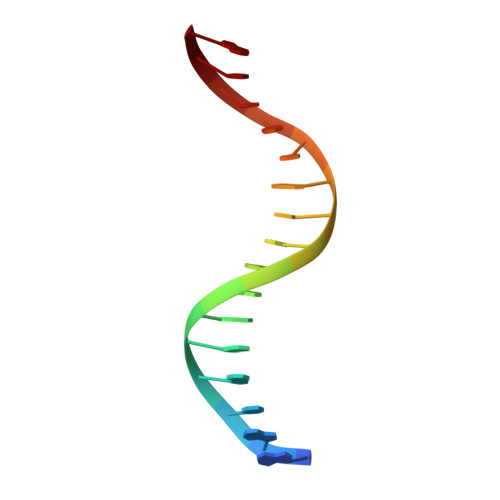
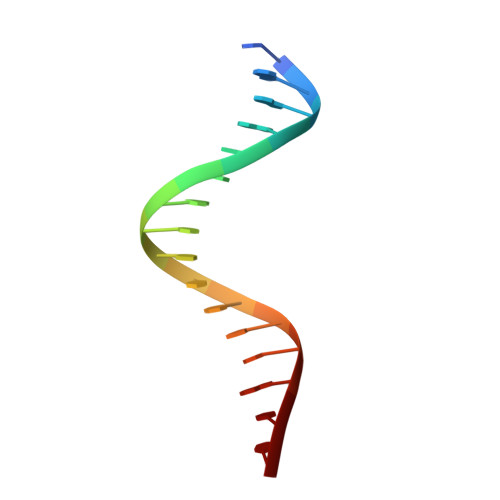
The SorC/DeoR family is a large family of bacterial transcription regulators that are involved in the control of carbohydrate metabolism and quorum sensing. To understand the structural basis of DNA recognition, structural studies of two functionally characterized SorC/DeoR family members from Bacillus subtilis were performed: the deoxyribonucleoside regulator bsDeoR and the central glycolytic genes regulator bsCggR. Each selected protein represents one of the subgroups that are recognized within the family. Crystal structures were determined of the N-terminal DNA-binding domains of bsDeoR and bsCggR in complex with DNA duplexes representing the minimal operator sequence at resolutions of 2.3 and 2.1 Å, respectively. While bsDeoR DBD contains a homeodomain-like HTH-type domain, bsCggR DBD contains a winged helix-turn-helix-type motif. Both proteins form C2-symmetric dimers that recognize two consecutive major grooves, and the protein-DNA interactions have been analyzed in detail. The crystal structures were used to model the interactions of the proteins with the full DNA operators, and a common mode of DNA recognition is proposed that is most likely to be shared by other members of the SorC/DeoR family.
- Structural Biology, Institute of Organic Chemistry and Biochemistry of Czech Academy of Sciences, Flemingovo nám. 2, 166 10 Prague, Czech Republic.
Organizational Affiliation: