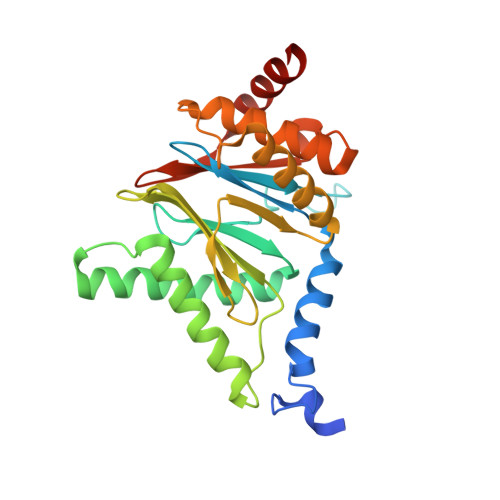
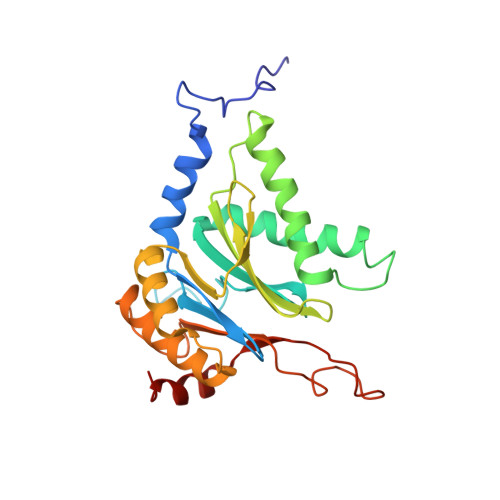
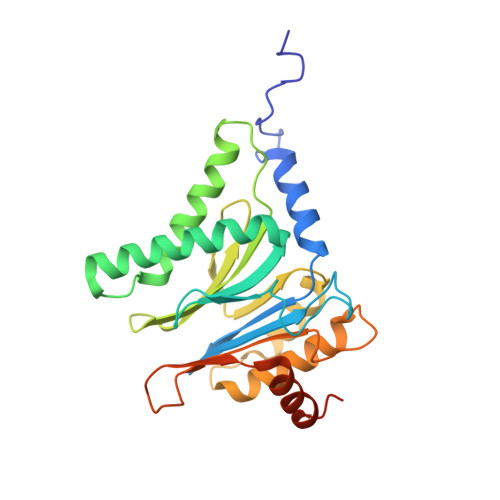
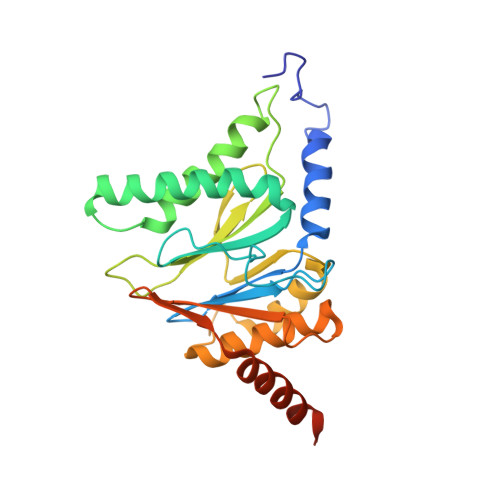
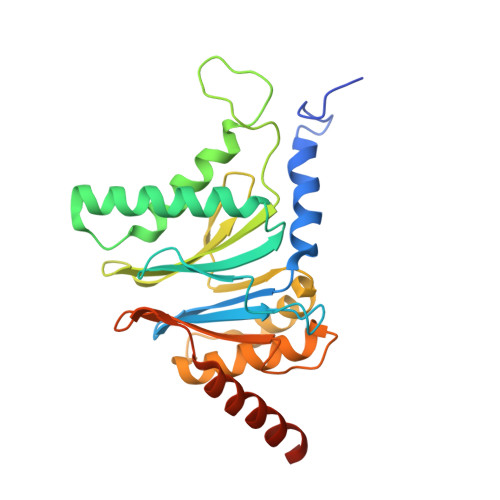
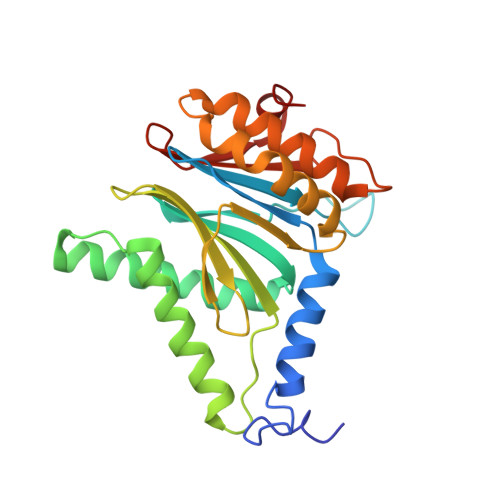
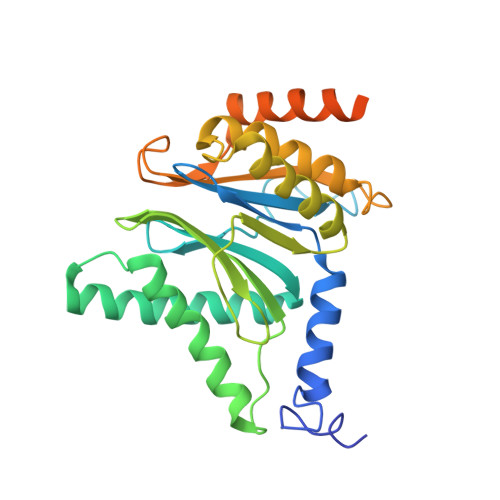
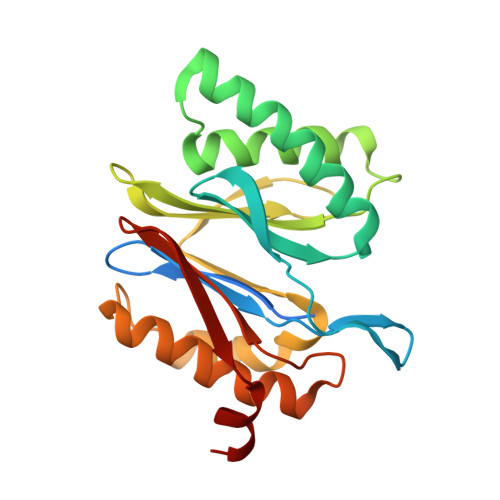
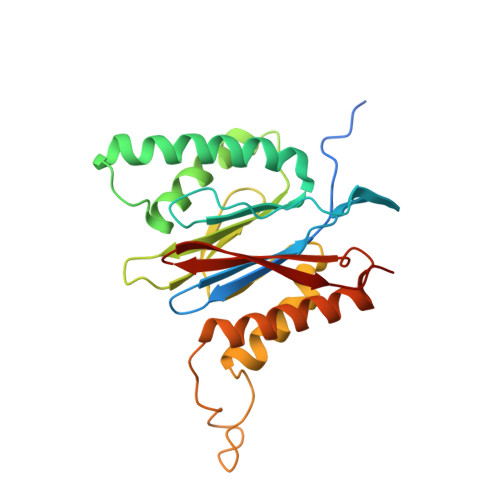
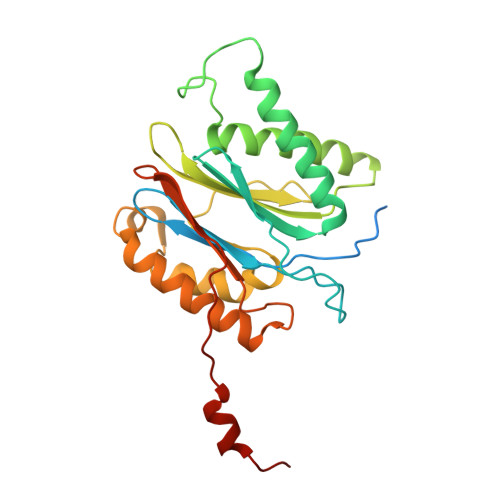
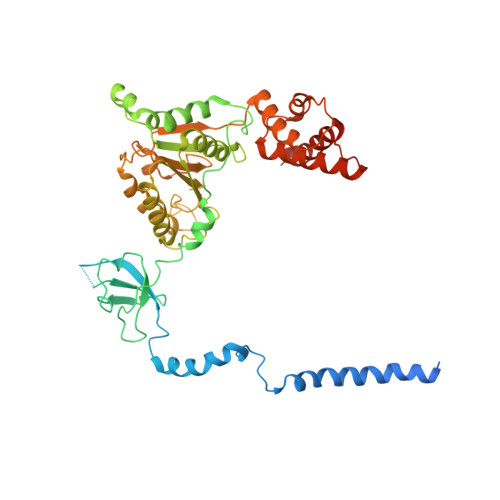
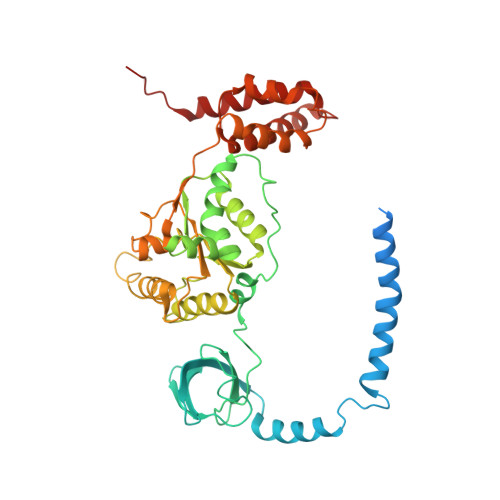
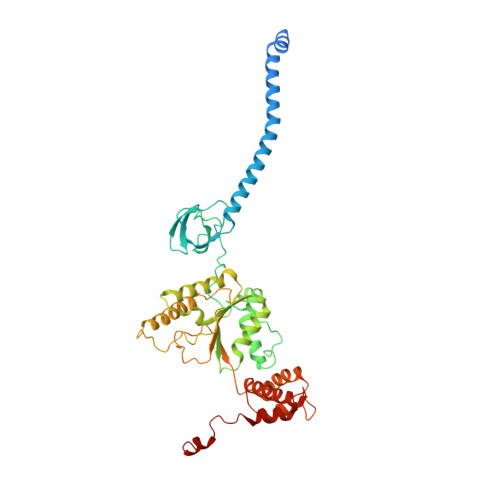
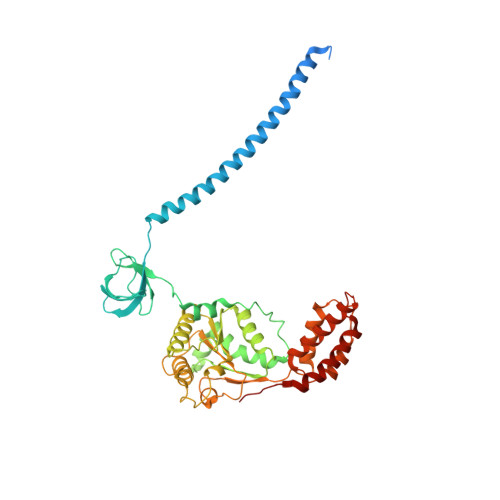
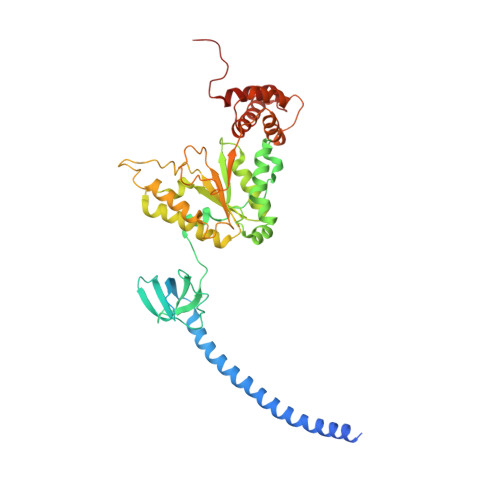
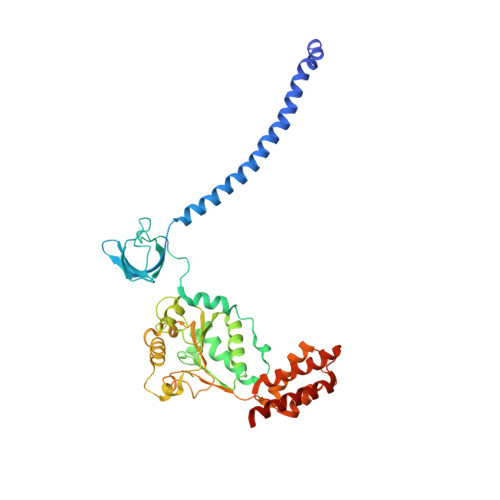
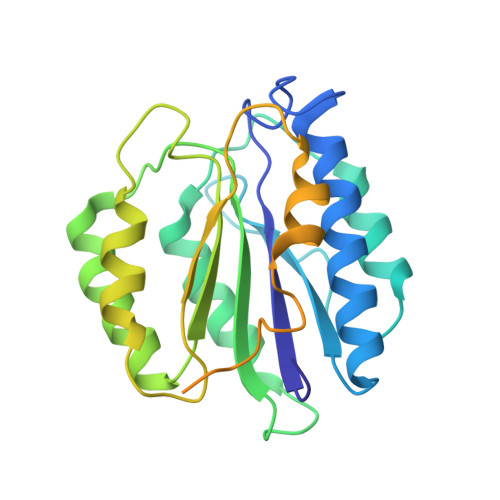
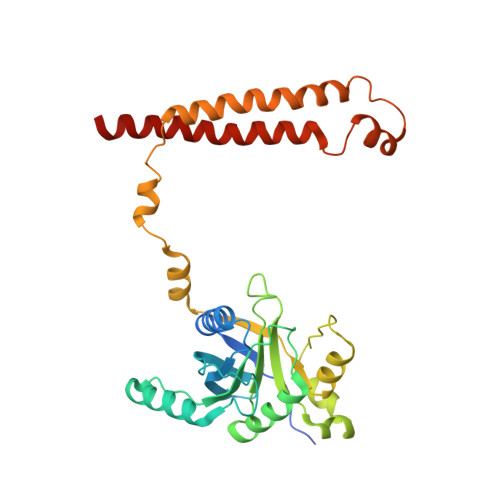
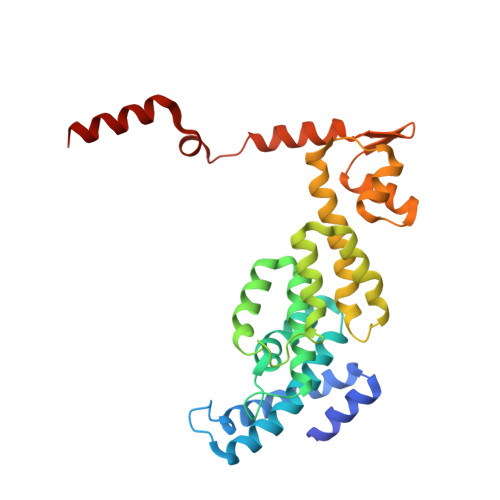
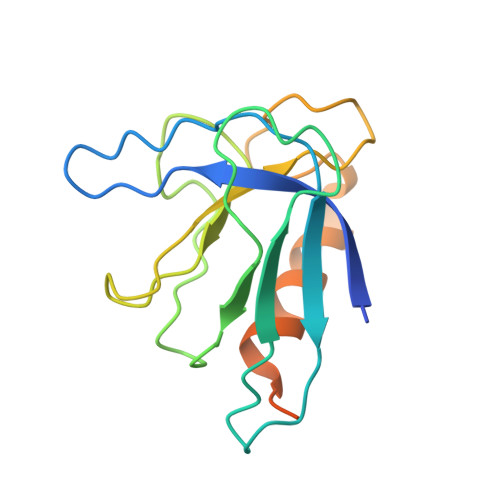
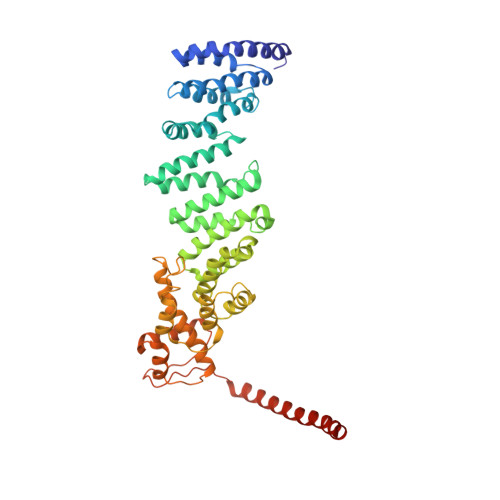
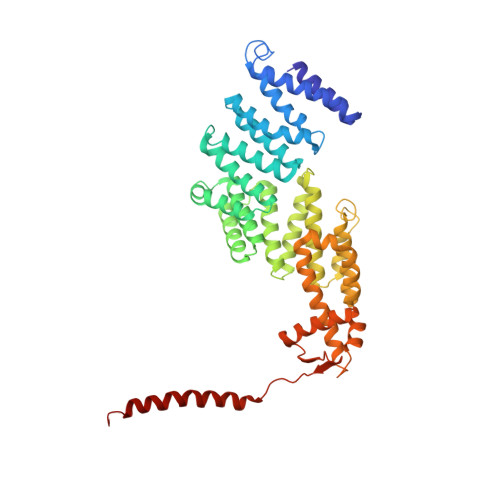
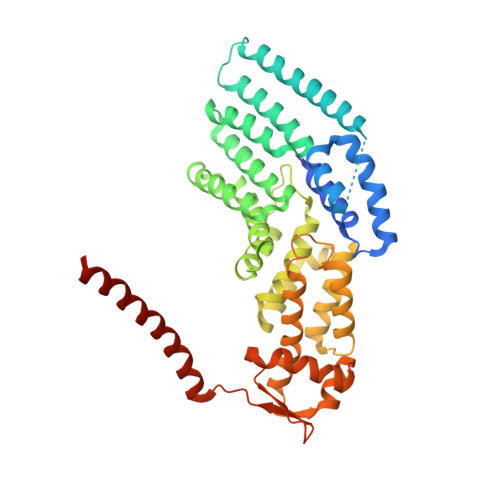
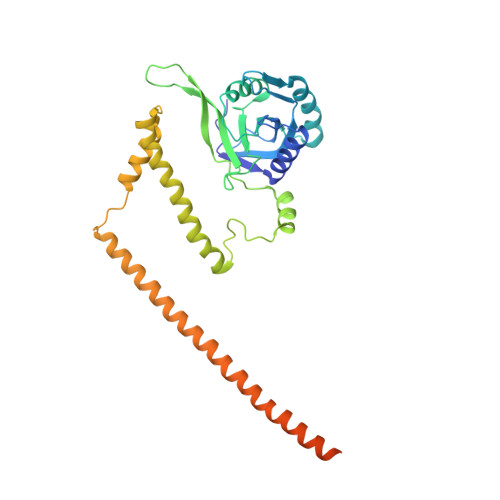
Structural insights into the functional cycle of the ATPase module of the 26S proteasome.
Wehmer, M., Rudack, T., Beck, F., Aufderheide, A., Pfeifer, G., Plitzko, J.M., Forster, F., Schulten, K., Baumeister, W., Sakata, E.(2017) Proc Natl Acad Sci U S A 114: 1305-1310
- PubMed: 28115689
- DOI: https://doi.org/10.1073/pnas.1621129114
- Primary Citation of Related Structures:
5MP9, 5MPA, 5MPB, 5MPC, 5MPD, 5MPE - PubMed Abstract:
In eukaryotic cells, the ubiquitin-proteasome system (UPS) is responsible for the regulated degradation of intracellular proteins. The 26S holocomplex comprises the core particle (CP), where proteolysis takes place, and one or two regulatory particles (RPs). The base of the RP is formed by a heterohexameric AAA + ATPase module, which unfolds and translocates substrates into the CP. Applying single-particle cryo-electron microscopy (cryo-EM) and image classification to samples in the presence of different nucleotides and nucleotide analogs, we were able to observe four distinct conformational states (s1 to s4). The resolution of the four conformers allowed for the construction of atomic models of the AAA + ATPase module as it progresses through the functional cycle. In a hitherto unobserved state (s4), the gate controlling access to the CP is open. The structures described in this study allow us to put forward a model for the 26S functional cycle driven by ATP hydrolysis.
- Department of Molecular Structural Biology, Max Planck Institute of Biochemistry, 82152 Martinsried, Germany.
Organizational Affiliation: