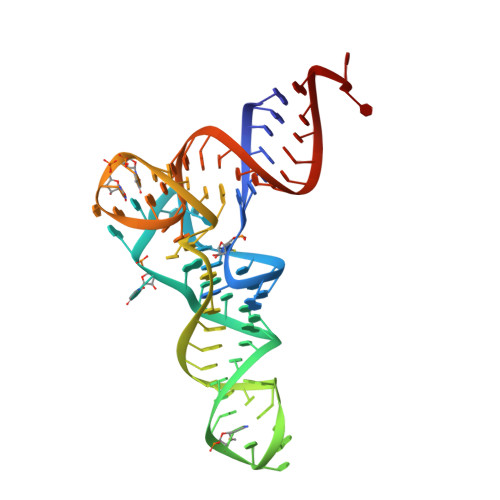
The structure of an E. coli tRNAf(Met) A1-U72 variant shows an unusual conformation of the A1-U72 base pair.
Monestier, A., Aleksandrov, A., Coureux, P.D., Panvert, M., Mechulam, Y., Schmitt, E.(2017) RNA 23: 673-682
- PubMed: 28143889
- DOI: https://doi.org/10.1261/rna.057877.116
- Primary Citation of Related Structures:
5L4O - PubMed Abstract:
Translation initiation in eukaryotes and archaea involves a methionylated initiator tRNA delivered to the ribosome in a ternary complex with e/aIF2 and GTP. Eukaryotic and archaeal initiator tRNAs contain a highly conserved A 1 -U 72 base pair at the top of the acceptor stem. The importance of this base pair to discriminate initiator tRNAs from elongator tRNAs has been established previously using genetics and biochemistry. However, no structural data illustrating how the A 1 -U 72 base pair participates in the accurate selection of the initiator tRNAs by the translation initiation systems are available. Here, we describe the crystal structure of a mutant E. coli initiator tRNA f Met A 1 -U 72 , aminoacylated with methionine, in which the C 1 :A 72 mismatch at the end of the tRNA acceptor stem has been changed to an A 1 -U 72 base pair. Sequence alignments show that the mutant E. coli tRNA is a good mimic of archaeal initiator tRNAs. The crystal structure, determined at 2.8 Å resolution, shows that the A 1 -U 72 pair adopts an unusual arrangement. A 1 is in a syn conformation and forms a single H-bond interaction with U 72 This interaction requires protonation of the N1 atom of A 1 Moreover, the 5' phosphoryl group folds back into the major groove of the acceptor stem and interacts with the N7 atom of G 2 A possible role of this unusual geometry of the A 1 -U 72 pair in the recognition of the initiator tRNA by its partners during eukaryotic and archaeal translation initiation is discussed.
- Laboratoire de Biochimie, Ecole polytechnique, CNRS, Université Paris-Saclay, 91128 Palaiseau cedex, France.
Organizational Affiliation: