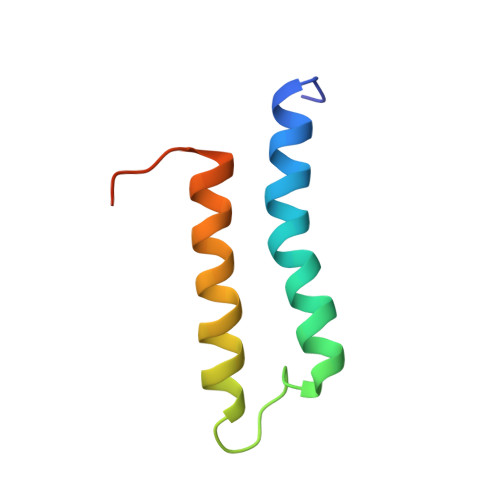
Solution structure of the microtubule-targeting COS domain of MID1.
Wright, K.M., Du, H., Dagnachew, M., Massiah, M.A.(2016) FEBS J 283: 3089-3102
- PubMed: 27367845
- DOI: https://doi.org/10.1111/febs.13795
- Primary Citation of Related Structures:
5IM8 - PubMed Abstract:
The human MID1 protein is required for the proper development during embryogenesis. Mutations of MID1 are associated with X-linked Opitz G syndrome, characterized by midline anomalies. MID1 associates with the microtubules and functions as an ubiquitin E3 ligase, targeting protein phosphatase 2A for ubiquitin-mediated regulation. The mechanism of microtubule association is not known. Recently, a 60-amino acid region termed the C-terminal subgroup One Signature (COS) box/domain was identified at the C-terminal end of the coiled-coil (CC) domain that facilitates microtubule localization. Insertion of the MID1 COS domain at the C-terminal end of the CC domain of a nonmicrotubule-associated TRIM protein confers microtubule localization. Here, we report the solution structure of the COS domain of MID1. The domain adopts a helix-loop-helix structure in which the N- and C-terminal ends are in close proximity. Hydrophobic residues stabilizing the interaction of the two α-helices form a central hydrophobic core. The loop separating the α-helices is structured, with two of its hydrophobic residues making contact with the central core. On the outer surface, positively charged residues form a distinct basic patch near the termini that we postulate is important for microtubule binding. A model of the structure of the preceding coiled-coil and COS domains (CC-COS) show that the COS domain forms a helical bundle at the C-terminal end of the CC domain similar to the spectrin-like fold observed with some known microtubule-binding proteins. Interestingly, the CC-COS domains bind to microtubules, demonstrating for the first time that MID1 can directly associate with the microtubules. Structural data are available in PDB database under the accession number 5IM8.
- Department of Chemistry and Center of Biomolecular Sciences, George Washington University, DC, USA.
Organizational Affiliation: