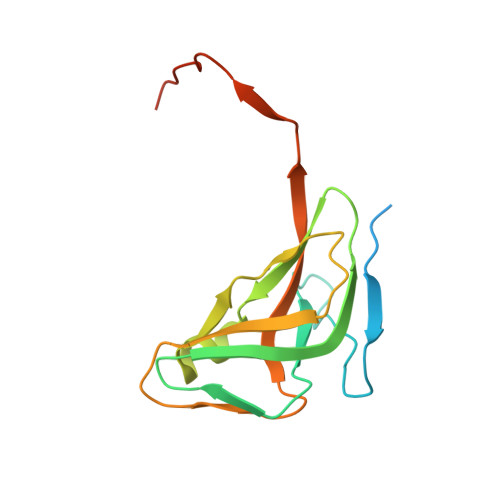
Structural insights into the mechanism defining substrate affinity in Arabidopsis thaliana dUTPase: the role of tryptophan 93 in ligand orientation.
Inoguchi, N., Chaiseeda, K., Yamanishi, M., Kim, M.K., Jang, Y., Bajaj, M., Chia, C.P., Becker, D.F., Moriyama, H.(2015) BMC Res Notes 8: 784-784
- PubMed: 26666293
- DOI: https://doi.org/10.1186/s13104-015-1760-1
- Primary Citation of Related Structures:
4OOP, 4OOQ - PubMed Abstract:
Deoxyuridine triphosphate nucleotidohydrolase (dUTPase) hydrolyzes dUTP to dUMP and pyrophosphate to maintain the cellular thymine-uracil ratio. dUTPase is also a target for cancer chemotherapy. However, the mechanism defining its substrate affinity remains unclear. Sequence comparisons of various dUTPases revealed that Arabidopsis thaliana dUTPase has a unique tryptophan at position 93, which potentially contributes to its degree of substrate affinity. To better understand the roles of tryptophan 93, A. thaliana dUTPase was studied. Enzyme assays showed that A. thaliana dUTPase belongs to a high-affinity group of isozymes, which also includes the enzymes from Escherichia coli and Mycobacterium tuberculosis. Enzymes from Homo sapiens and Saccharomyces cerevisiae are grouped as low-affinity dUTPases. The structure of the homo-trimeric A. thaliana dUTPase showed three active sites, each with a different set of ligand interactions between the amino acids and water molecules. On an α-helix, tryptophan 93 appears to keep serine 89 in place via a water molecule and to specifically direct the ligand. Upon being oriented in the active site, the C-terminal residues close the active site to promote the reaction. In the high-affinity group, the prefixed direction of the serine residues was oriented by a positively charged residue located four amino acids away, while low-affinity enzymes possess small hydrophobic residues at the corresponding sites.
- School of Biological Sciences, University of Nebraska-Lincoln, Lincoln, NE, USA. ninoguchi@huskers.unl.edu.
Organizational Affiliation: