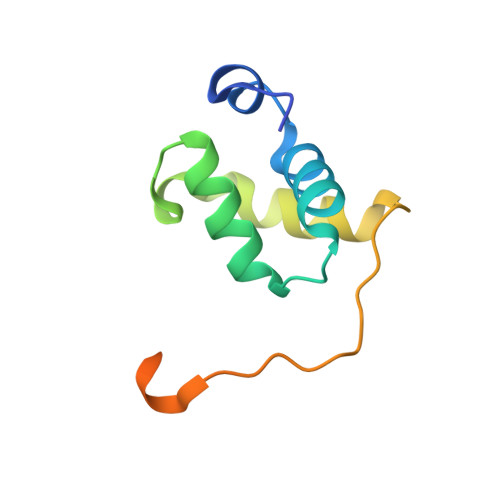
X-ray vs. NMR structure of N-terminal domain of delta-subunit of RNA polymerase.
Demo, G., Papouskova, V., Komarek, J., Kaderavek, P., Otrusinova, O., Srb, P., Rabatinova, A., Krasny, L., Zidek, L., Sklenar, V., Wimmerova, M.(2014) J Struct Biol 187: 174-186
- PubMed: 24937760
- DOI: https://doi.org/10.1016/j.jsb.2014.06.001
- Primary Citation of Related Structures:
4NC7, 4NC8 - PubMed Abstract:
The crystal structure of the N-terminal domain of the RNA polymerase δ subunit (Nδ) from Bacillus subtilis solved at a resolution of 2.0Å is compared with the NMR structure determined previously. The molecule crystallizes in the space group C222(1) with a dimer in the asymmetric unit. Importantly, the X-ray structure exhibits significant differences from the lowest energy NMR structure. In addition to the overall structure differences, structurally important β sheets found in the NMR structure are not present in the crystal structure. We systematically investigated the cause of the discrepancies between the NMR and X-ray structures of Nδ, addressing the pH dependence, presence of metal ions, and crystal packing forces. We convincingly showed that the crystal packing forces, together with the presence of Ni(2+) ions, are the main reason for such a difference. In summary, the study illustrates that the two structural approaches may give unequal results, which need to be interpreted with care to obtain reliable structural information in terms of biological relevance.
- National Centre for Biomolecular Research, Faculty of Science, Masaryk University, Kamenice 5/A4, 62500 Brno, Czech Republic; Central European Institute of Technology-CEITEC, Masaryk University, Kamenice 5/A4, 62500 Brno, Czech Republic.
Organizational Affiliation: