Specificity and Structure of a High Affinity Activin Receptor-like Kinase 1 (ALK1) Signaling Complex.
Townson, S.A., Martinez-Hackert, E., Greppi, C., Lowden, P., Sako, D., Liu, J., Ucran, J.A., Liharska, K., Underwood, K.W., Seehra, J., Kumar, R., Grinberg, A.V.(2012) J Biological Chem 287: 27313-27325
- PubMed: 22718755
- DOI: https://doi.org/10.1074/jbc.M112.377960
- Primary Citation of Related Structures:
4FAO - PubMed Abstract:
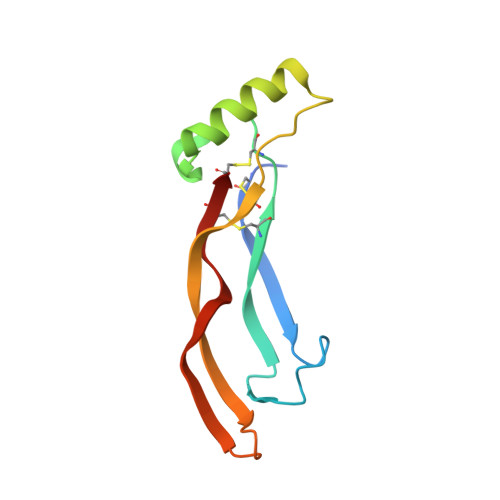
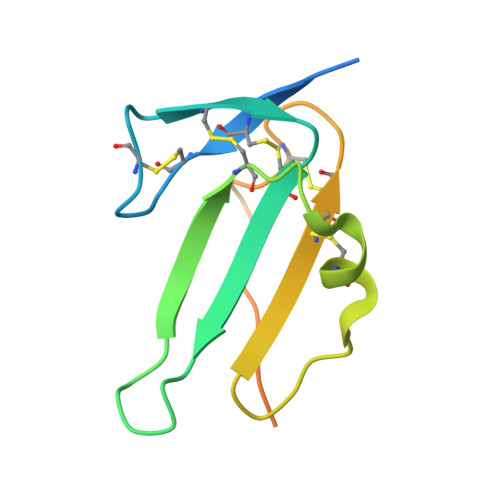
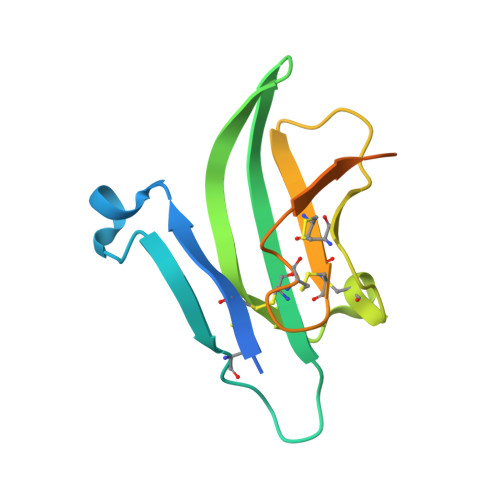
Activin receptor-like kinase 1 (ALK1), an endothelial cell-specific type I receptor of the TGF-β superfamily, is an important regulator of normal blood vessel development as well as pathological tumor angiogenesis. As such, ALK1 is an important therapeutic target. Thus, several ALK1-directed agents are currently in clinical trials as anti-angiogenic cancer therapeutics. Given the biological and clinical importance of the ALK1 signaling pathway, we sought to elucidate the biophysical and structural basis underlying ALK1 signaling. The TGF-β family ligands BMP9 and BMP10 as well as the three type II TGF-β family receptors ActRIIA, ActRIIB, and BMPRII have been implicated in ALK1 signaling. Here, we provide a kinetic and thermodynamic analysis of BMP9 and BMP10 interactions with ALK1 and type II receptors. Our data show that BMP9 displays a significant discrimination in type II receptor binding, whereas BMP10 does not. We also report the crystal structure of a fully assembled ternary complex of BMP9 with the extracellular domains of ALK1 and ActRIIB. The structure reveals that the high specificity of ALK1 for BMP9/10 is determined by a novel orientation of ALK1 with respect to BMP9, which leads to a unique set of receptor-ligand interactions. In addition, the structure explains how BMP9 discriminates between low and high affinity type II receptors. Taken together, our findings provide structural and mechanistic insights into ALK1 signaling that could serve as a basis for novel anti-angiogenic therapies.
- Acceleron Pharma, Cambridge, Massachusetts 02139, USA.
Organizational Affiliation: