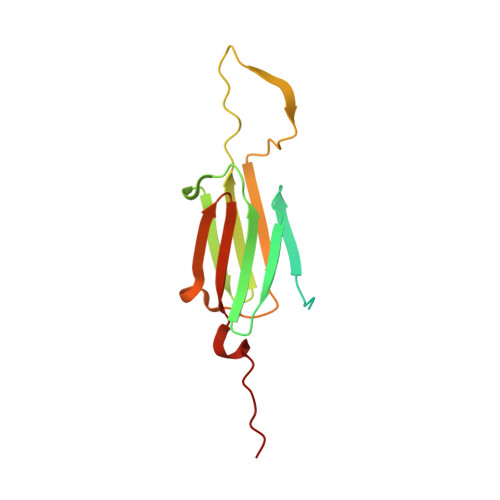
Crystal structure of an activated variant of small heat shock protein hsp16.5.
McHaourab, H.S., Lin, Y.L., Spiller, B.W.(2012) Biochemistry 51: 5105-5112
- PubMed: 22670769
- DOI: https://doi.org/10.1021/bi300525x
- Primary Citation of Related Structures:
4ELD - PubMed Abstract:
How does the sequence of a single small heat shock protein (sHSP) assemble into oligomers of different sizes? To gain insight into the underlying structural mechanism, we determined the crystal structure of an engineered variant of Methanocaldococcus jannaschii Hsp16.5 wherein a 14 amino acid peptide from human heat shock protein 27 (Hsp27) was inserted at the junction of the N-terminal region and the α-crystallin domain. In response to this insertion, the oligomer shell expands from 24 to 48 subunits while maintaining octahedral symmetry. Oligomer rearrangement does not alter the fold of the conserved α-crystallin domain nor does it disturb the interface holding the dimeric building block together. Rather, the flexible C-terminal tail of Hsp16.5 changes its orientation relative to the α-crystallin domain which enables alternative packing of dimers. This change in orientation preserves a peptide-in-groove interaction of the C-terminal tail with an adjacent β-sandwich, thereby holding the assembly together. The interior of the expanded oligomer, where substrates presumably bind, retains its predominantly nonpolar character relative to the outside surface. New large windows in the outer shell provide increased access to these substrate-binding regions, thus accounting for the higher affinity of this variant to substrates. Oligomer polydispersity regulates sHSPs chaperone activity in vitro and has been implicated in their physiological roles. The structural mechanism of Hsp16.5 oligomer flexibility revealed here, which is likely to be highly conserved across the sHSP superfamily, explains the relationship between oligomer expansion observed in disease-linked mutants and changes in chaperone activity.
- Department of Molecular Physiology and Biophysics, Vanderbilt University Medical Center, Tennessee 37232, USA. hassane.mchaourab@vanderbilt.edu
Organizational Affiliation: