Resolution of disulfide heterogeneity in Nogo receptor 1 fusion proteins by molecular engineering.
Weinreb, P.H., Wen, D., Qian, F., Wildes, C.P., Garber, E.A., Walus, L., Jung, M.Y., Wang, J., Relton, J.K., Amatucci, J., Wang, R., Porreca, F., Silvian, L., Meier, W., Pepinsky, R.B., Lee, D.H.(2010) Biotechnol Appl Biochem 57: 31-45
- PubMed: 20815818
- DOI: https://doi.org/10.1042/BA20100061
- Primary Citation of Related Structures:
3KJ4 - PubMed Abstract:
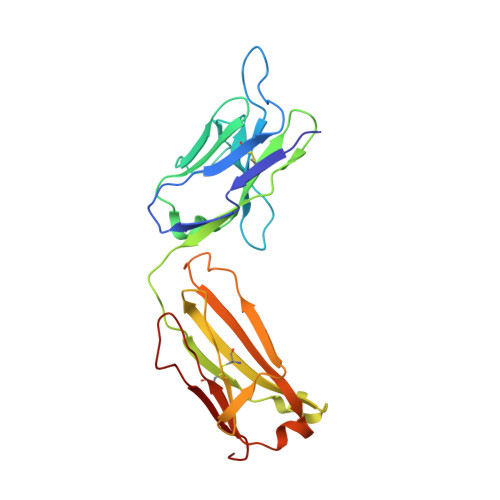
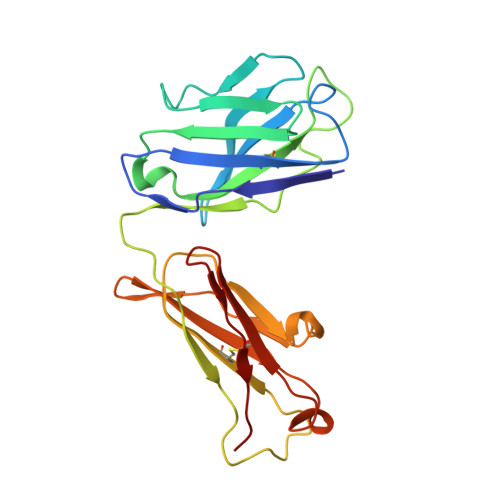
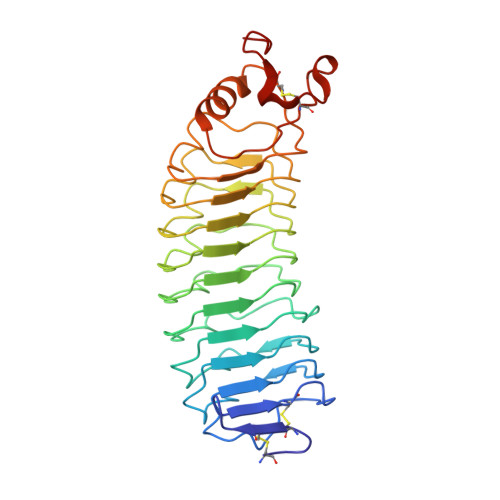
NgRI (Nogo-66 receptor) is part of a signalling complex that inhibits axon regeneration in the central nervous system. Truncated soluble versions of NgRI have been used successfully to promote axon regeneration in animal models of spinal-cord injury, raising interest in this protein as a potential therapeutic target. The LRR (leucine-rich repeat) regions in NgRI are flanked by N- and C-terminal disulfide-containing 'cap' domains (LRRNT and LRRCT respectively). In the present work we show that, although functionally active, the NgRI(310)-Fc fusion protein contains mislinked and heterogeneous disulfide patterns in the LRRCT domain, and we report the generation of a series of variant molecules specifically designed to prevent this heterogeneity. Using these variants we explored the effects of modifying the NgRI truncation site or the spacing between the NgRI and Fc domains, or replacing cysteines within the NgRI or IgG hinge regions. One variant, which incorporates replacements of Cys²⁶⁶ and Cys³⁰⁹ with alanine residues, completely eliminated disulfide scrambling while maintaining functional in vitro and in vivo efficacy. This modified NgRI-Fc molecule represents a significantly improved candidate for further pharmaceutical development, and may serve as a useful model for the optimization of other IgG fusion proteins made from LRR proteins.
- Biogen Idec, Inc., 14 Cambridge Center, Cambridge, MA 02142, USA. paul.weinreb@biogenidec.com
Organizational Affiliation: