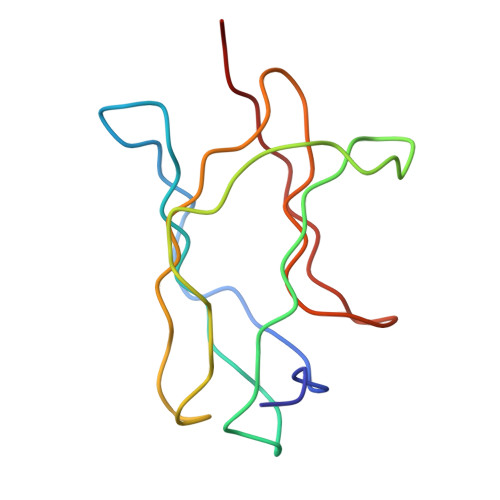
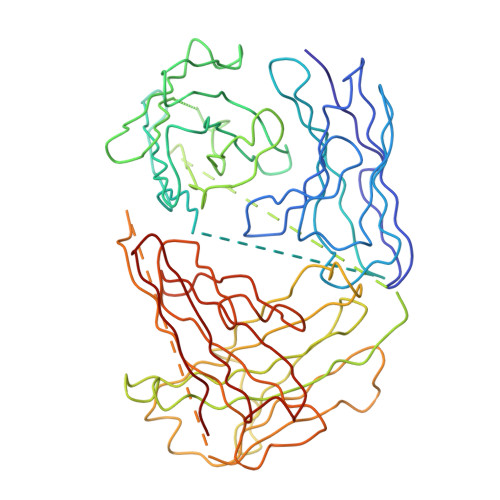
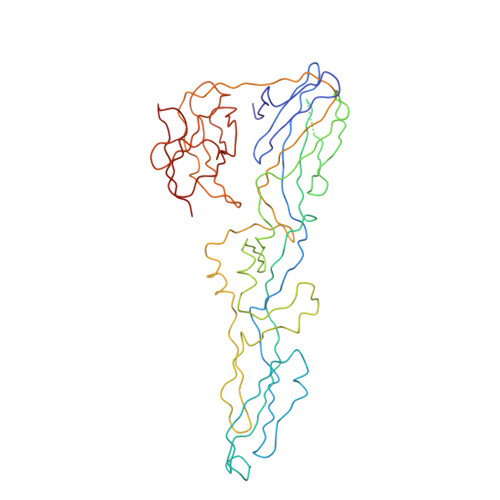
Structure of Acidic pH Dengue Virus Showing the Fusogenic Glycoprotein Trimers.
Zhang, X., Sheng, J., Austin, S.K., Hoornweg, T.E., Smit, J.M., Kuhn, R.J., Diamond, M.S., Rossmann, M.G.(2015) J Virol 89: 743-750
- PubMed: 25355881
- DOI: https://doi.org/10.1128/JVI.02411-14
- Primary Citation of Related Structures:
3J8D - PubMed Abstract:
Flaviviruses undergo large conformational changes during their life cycle. Under acidic pH conditions, the mature virus forms transient fusogenic trimers of E glycoproteins that engage the lipid membrane in host cells to initiate viral fusion and nucleocapsid penetration into the cytoplasm. However, the dynamic nature of the fusogenic trimer has made the determination of its structure a challenge. Here we have used Fab fragments of the neutralizing antibody DV2-E104 to stop the conformational change of dengue virus at an intermediate stage of the fusion process. Using cryo-electron microscopy, we show that in this intermediate stage, the E glycoproteins form 60 trimers that are similar to the predicted "open" fusogenic trimer. The structure of a dengue virus has been captured during the formation of fusogenic trimers. This was accomplished by binding Fab fragments of the neutralizing antibody DV2-E104 to the virus at neutral pH and then decreasing the pH to 5.5. These trimers had an "open" conformation, which is distinct from the "closed" conformation of postfusion trimers. Only two of the three E proteins within each spike are bound by a Fab molecule at domain III. Steric hindrance around the icosahedral 3-fold axes prevents binding of a Fab to the third domain III of each E protein spike. Binding of the DV2-E104 Fab fragments prevents domain III from rotating by about 130° to the postfusion orientation and thus precludes the stem region from "zipping" together the three E proteins along the domain II boundaries into the "closed" postfusion conformation, thus inhibiting fusion.
- Department of Biological Sciences, Purdue University, West Lafayette, Indiana, USA.
Organizational Affiliation: