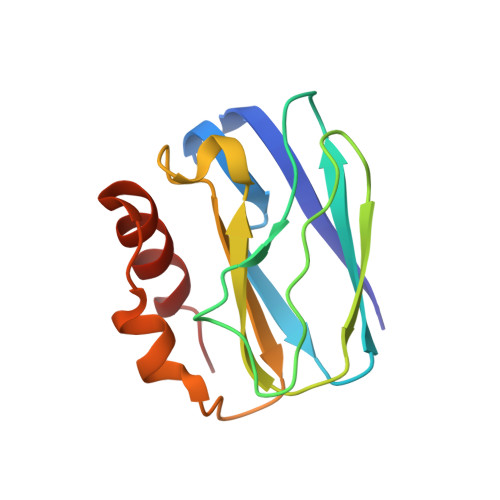
The 1.4 A resolution structure of Paracoccus pantotrophus pseudoazurin.
Najmudin, S., Pauleta, S.R., Moura, I., Romao, M.J.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 627-635
- PubMed: 20516588
- DOI: https://doi.org/10.1107/S1744309110013989
- Primary Citation of Related Structures:
3ERX - PubMed Abstract:
Pseudoazurins are small type 1 copper proteins that are involved in the flow of electrons between various electron donors and acceptors in the bacterial periplasm, mostly under denitrifying conditions. The previously determined structure of Paracoccus pantotrophus pseudoazurin in the oxidized form was improved to a nominal resolution of 1.4 A, with R and R(free) values of 0.188 and 0.206, respectively. This high-resolution structure makes it possible to analyze the interactions between the monomers and the solvent structure in detail. Analysis of the high-resolution structure revealed the structural regions that are responsible for monomer-monomer recognition during dimer formation and for protein-protein interaction and that are important for partner recognition. The pseudoazurin structure was compared with other structures of various type 1 copper proteins and these were grouped into families according to similarities in their secondary structure; this may be useful in the annotation of copper proteins in newly sequenced genomes and in the identification of novel copper proteins.
- REQUIMTE, Centro de Química Fina e Biotecnologia, Departamento de Química, Faculdade de Ciências e Tecnologia, Universidade Nova de Lisboa, 2829-516 Caparica, Portugal. shabir@fmv.utl.pt
Organizational Affiliation: