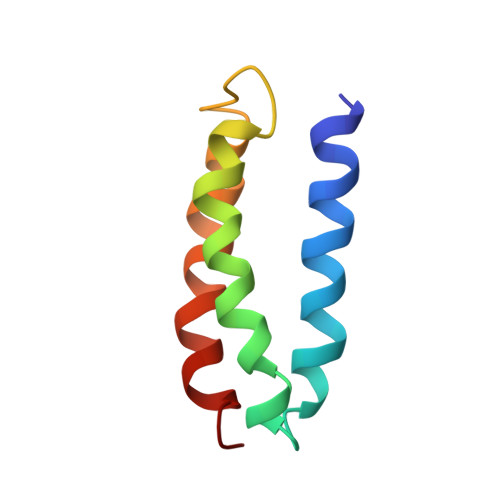
Solution structure and dynamics of a de novo designed three-helix bundle protein.
Walsh, S.T., Cheng, H., Bryson, J.W., Roder, H., DeGrado, W.F.(1999) Proc Natl Acad Sci U S A 96: 5486-5491
- PubMed: 10318910
- DOI: https://doi.org/10.1073/pnas.96.10.5486
- Primary Citation of Related Structures:
2A3D - PubMed Abstract:
Although de novo protein design is an important endeavor with implications for understanding protein folding, until now, structures have been determined for only a few 25- to 30-residue designed miniproteins. Here, the NMR solution structure of a complex 73-residue three-helix bundle protein, alpha3D, is reported. The structure of alpha3D was not based on any natural protein, and yet it shows thermodynamic and spectroscopic properties typical of native proteins. A variety of features contribute to its unique structure, including electrostatics, the packing of a diverse set of hydrophobic side chains, and a loop that incorporates common capping motifs. Thus, it is now possible to design a complex protein with a well defined and predictable three-dimensional structure.
- The Johnson Research Foundation, Department of Biochemistry and Biophysics, University of Pennsylvania, Philadelphia, PA 19104, USA.
Organizational Affiliation: