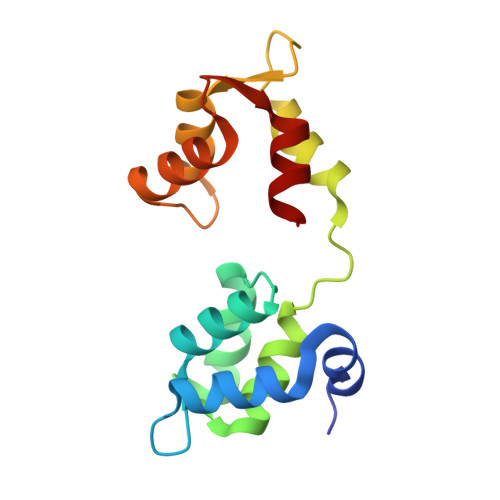
The Closed MTIP-Myosin A-Tail Complex from the Malaria Parasite Invasion Machinery.
Bosch, J., Turley, S., Roach, C.M., Daly, T.M., Bergman, L.W., Hol, W.G.(2007) J Mol Biology 372: 77-88
- PubMed: 17628590
- DOI: https://doi.org/10.1016/j.jmb.2007.06.016
- Primary Citation of Related Structures:
2QAC - PubMed Abstract:
The Myosin A-tail interacting protein (MTIP) of the malaria parasite links the actomyosin motor of the host cell invasion machinery to its inner membrane complex. We report here that at neutral pH Plasmodium falciparum MTIP in complex with Myosin A adopts a compact conformation, with its two domains completely surrounding the Myosin A-tail helix, dramatically different from previously observed extended MTIP structures. Crystallographic and mutagenesis studies show that H810 and K813 of Myosin A are key players in the formation of the compact MTIP:Myosin A complex. Only the unprotonated state of Myosin A-H810 is compatible with the compact complex. Most surprisingly, every side-chain atom of Myosin A-K813 is engaged in contacts with MTIP. While this side-chain was previously considered to prevent a compact conformation of MTIP with Myosin A, it actually appears to be essential for the formation of the compact complex. The hydrophobic pockets and adaptability seen in the available series of MTIP structures bodes well for the discovery of inhibitors of cell invasion by malaria parasites.
- Department of Biochemistry, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: