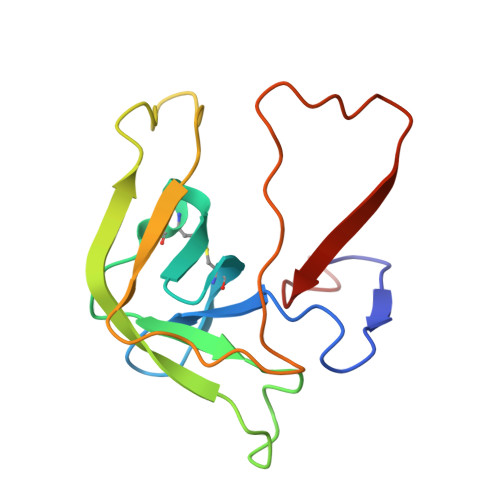
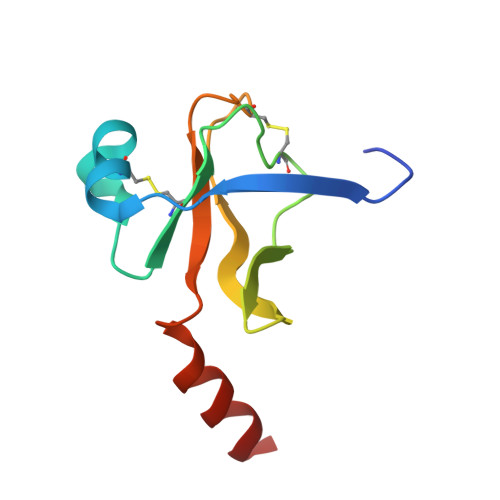
Inhibition of chymotrypsin by a complex of ortho-vanadate and benzohydroxamic Acid: structure of the inert complex and its mechanistic interpretation.
Moulin, A., Bell, J.H., Pratt, R.F., Ringe, D.(2007) Biochemistry 46: 5982-5990
- PubMed: 17469803
- DOI: https://doi.org/10.1021/bi6025209
- Primary Citation of Related Structures:
2P8O - PubMed Abstract:
Serine proteases, like serine beta-lactamases, are rapidly and covalently inhibited by suitably designed phosph(on)ates. The active sites of these enzymes must, therefore, be able to stabilize the pentacoordinated transition states of phosphyl transfer reactions as well as the tetrahedral transition states of acyl transfers. It follows that these enzymes should also be inhibited by molecules capable of generating inert pentacoordinated species. We (J.H.B. and R.F.P.) have previously shown that these enzymes are, in fact, rapidly and reversibly inhibited by 1:1 complexes of vanadate and hydroxamic acids. In this paper, we present the first crystal structure of an acyl transferase inhibited by vanadate. The complex of vanadate and benzohydroxamic acid is a competitive inhibitor of alpha-chymotrypsin with a KI value of 16 muM. In the structure, obtained at a resolution of 1.5 A, the protein is conformationally little different from the apoenzyme. The vanadium, in a distorted octahedral ligand field, is covalently bound to the active site serine oxygen group. One oxgen ligand, presumably anionic, is located in the oxyanion hole. Another is directed roughly in the direction of the acyl transfer leaving group, and a third in the direction of the S2 site. The hydroxamate is bound to vanadium through the hydroxyl oxygen and also, more weakly, through the carbonyl group, to form a five-membered chelate ring. The effect of this chelation is to place the phenyl group of the inhibitor into the important S1 specificity site. The hydroxamate oxygen is directed in line away from the Ser195 Ogamma, approximating the direction of departure of a leaving group in phosphyl transfer. The entire complex can be seen as a reasonable mimic of a phosphyl transfer transition state where the leaving group is extended into the S1 site.
- Rosenstiel Basic Medical Sciences Research Center, Program in Biochemistry, and Program in Biophysics, Brandeis University, Waltham, Massachusetts 02454, USA.
Organizational Affiliation: