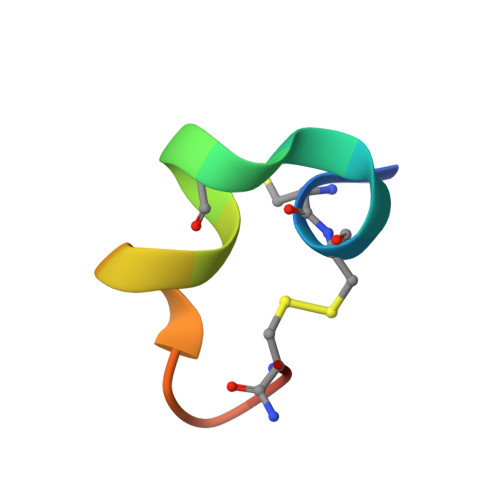
A novel alpha 4/7-conotoxin LvIA from Conus lividus that selectively blocks alpha 3 beta 2 vs. alpha 6/ alpha 3 beta 2 beta 3 nicotinic acetylcholine receptors.
Luo, S., Zhangsun, D., Schroeder, C.I., Zhu, X., Hu, Y., Wu, Y., Weltzin, M.M., Eberhard, S., Kaas, Q., Craik, D.J., McIntosh, J.M., Whiteaker, P.(2014) FASEB J 28: 1842-1853
- PubMed: 24398291
- DOI: https://doi.org/10.1096/fj.13-244103
- Primary Citation of Related Structures:
2MDQ - PubMed Abstract:
This study was performed to discover and characterize the first potent α3β2-subtype-selective nicotinic acetylcholine receptor (nAChR) ligand. A novel α4/7-conotoxin, α-CTxLvIA, was cloned from Conus lividus. Its pharmacological profile at Xenopus laevis oocyte-expressed rat nAChR subtypes was determined by 2-electrode voltage-clamp electrophysiology, and its 3-dimensional (3D) structure was determined by NMR spectroscopy. α-CTx LvIA is a 16-aa C-terminally-amidated peptide with 2-disulfide bridges. Using rat subunits expressed in Xenopus oocytes, we found the highest affinity of α-CTxLvIA was for α3β2 nAChRs (IC50 8.7 nM), where blockade was reversible within 2 min. IC50 values were >100 nM at α6/α3β2β3, α6/α3β4, and α3β4 nAChRs, and ≥3 μM at all other subtypes tested. α3β2 vs. α6β2 subtype selectivity was confirmed for human-subunit nAChRs with much greater preference (300-fold) for α3β2 over α6β2 nAChRs. This is the first α-CTx reported to show high selectivity for human α3β2 vs. α6β2 nAChRs. α-CTxLvIA adopts two similarly populated conformations water: one (assumed to be bioactive) is highly structured, whereas the other is mostly random coil in nature. Selectivity differences with the similarly potent, but less selective, α3β2 nAChR antagonist α-CTx PeIA probably reside within the three residues, which differ in loop 2, given their otherwise similar 3D structures. α4/7-CTx LvIA is a new, potent, selective α3β2 nAChR antagonist, which will enable detailed studies of α3β2 nAChR structure, function, and physiological roles.
- 1Key Laboratory of Tropical Biological Resources, Ministry of Education, Hainan University; Haikou, Hainan, 570228 China. luosulan2003@163.com.
Organizational Affiliation: