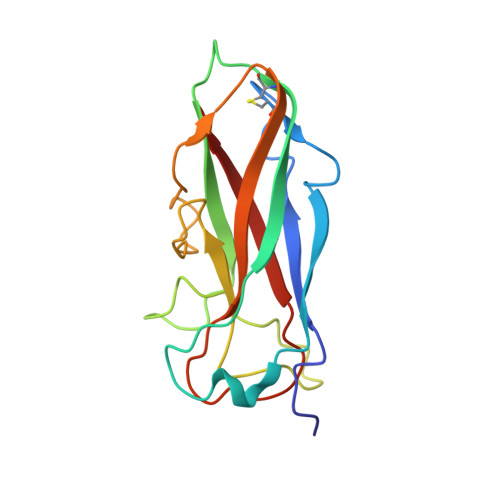
Intramolecular donor strand complementation in the E. coli type 1 pilus subunit FimA explains the existence of FimA monomers as off-pathway products of pilus assembly that inhibit host cell apoptosis.
Walczak, M.J., Puorger, C., Glockshuber, R., Wider, G.(2014) J Mol Biology 426: 542-549
- PubMed: 24184277
- DOI: https://doi.org/10.1016/j.jmb.2013.10.029
- Primary Citation of Related Structures:
2M5G - PubMed Abstract:
Type 1 pili are filamentous organelles mediating the attachment of uropathogenic Escherichia coli to epithelial cells of host organisms. The helical pilus rod consists of up to 3000 copies of the main structural subunit FimA that interact via donor strand complementation, where the incomplete Ig-like fold of FimA is completed by insertion of the N-terminal extension (donor strand) of the following FimA subunit. Recently, it was shown that FimA also exists in a monomeric, assembly-incompetent form and that FimA monomers act as inhibitors of apoptosis in infected host cells. Here we present the NMR structure of monomeric wild-type FimA with its natural N-terminal donor strand complementing the Ig fold. Compared to FimA subunits in the assembled pilus, intramolecular self-complementation in the monomer stabilizes the FimA fold with significantly less interactions, and the natural FimA donor strand is inserted in the opposite orientation. In addition, we show that a motif of two glycine residues in the FimA donor strand, separated by five residues, is the prerequisite of the alternative, parallel donor strand insertion mechanism in the FimA monomer and that this motif is preserved in FimA homologs of many enteroinvasive pathogens. We conclude that FimA is a unique case of a protein with alternative, functionally relevant folding possibilities, with the FimA polymer forming the highly stable pilus rod and the FimA monomer promoting pathogen propagation by apoptosis suppression of infected epithelial target cells.
- ETH Zurich Institute of Molecular Biology and Biophysics, CH-8093 Zurich, Switzerland.
Organizational Affiliation: