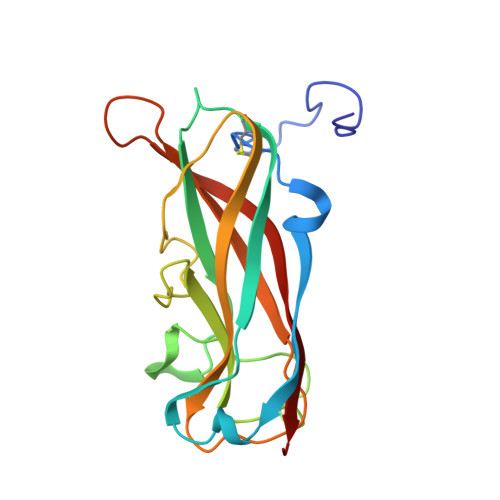
Structure, Folding and Stability of FimA, the Main Structural Subunit of Type 1 Pili from Uropathogenic Escherichia coli Strains.
Puorger, C., Vetsch, M., Wider, G., Glockshuber, R.(2011) J Mol Biology 412: 520-535
- PubMed: 21816158
- DOI: https://doi.org/10.1016/j.jmb.2011.07.044
- Primary Citation of Related Structures:
2JTY - PubMed Abstract:
Filamentous type 1 pili are responsible for attachment of uropathogenic Escherichia coli strains to host cells. They consist of a linear tip fibrillum and a helical rod formed by up to 3000 copies of the main structural pilus subunit FimA. The subunits in the pilus interact via donor strand complementation, where the incomplete, immunoglobulin-like fold of each subunit is complemented by an N-terminal donor strand of the subsequent subunit. Here, we show that folding of FimA occurs at an extremely slow rate (half-life: 1.6 h) and is catalyzed more than 400-fold by the pilus chaperone FimC. Moreover, FimA is capable of intramolecular self-complementation via its own donor strand, as evidenced by the loss of folding competence upon donor strand deletion. Folded FimA is an assembly-incompetent monomer of low thermodynamic stability (-10.1 kJ mol(-1)) that can be rescued for pilus assembly at 37 °C because FimC selectively pulls the fraction of unfolded FimA molecules from the FimA folding equilibrium and allows FimA refolding on its surface. Elongation of FimA at the C-terminus by its own donor strand generated a self-complemented variant (FimAa) with alternative folding possibilities that spontaneously adopts the more stable conformation (-85.0 kJ mol(-1)) in which the C-terminal donor strand is inserted in the opposite orientation relative to that in FimA. The solved NMR structure of FimAa revealed extensive β-sheet hydrogen bonding between the FimA pilin domain and the C-terminal donor strand and provides the basis for reconstruction of an atomic model of the pilus rod.
- ETH Zürich, Institute of Molecular Biology and Biophysics, 8093 Zurich, Switzerland.
Organizational Affiliation: