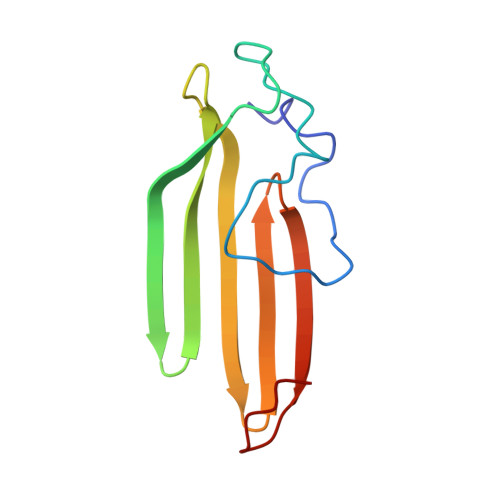
Embryonic Neural Inducing Factor Churchill Is not a DNA-binding Zinc Finger Protein: Solution Structure Reveals a Solvent-exposed beta-Sheet and Zinc Binuclear Cluster
Lee, B.M., Buck-Koehntop, B.A., Martinez-Yamout, M.A., Dyson, H.J., Wright, P.E.(2007) J Mol Biology 371: 1274-1289
- PubMed: 17610897
- DOI: https://doi.org/10.1016/j.jmb.2007.06.021
- Primary Citation of Related Structures:
2JOX - PubMed Abstract:
Churchill is a zinc-containing protein that is involved in neural induction during embryogenesis. At the time of its discovery, it was thought on the basis of sequence alignment to contain two zinc fingers of the C4 type. Further, binding of an N-terminal GST-Churchill fusion protein to a particular DNA sequence was demonstrated by immunoprecipitation selection assay, suggesting that Churchill may function as a transcriptional regulator by sequence-specific DNA binding. We show by NMR solution structure determination that, far from containing canonical C4 zinc fingers, the protein contains three bound zinc ions in novel coordination sites, including an unusual binuclear zinc cluster. The secondary structure of Churchill is also unusual, consisting of a highly solvent-exposed single-layer beta-sheet. Hydrogen-deuterium exchange and backbone relaxation measurements reveal that Churchill is unusually dynamic on a number of time scales, with the exception of regions surrounding the zinc coordinating sites, which serve to stabilize the otherwise unstructured N terminus and the single-layer beta-sheet. No binding of Churchill to the previously identified DNA sequence could be detected, and extensive searches using DNA sequence selection techniques could find no other DNA sequence that was bound by Churchill. Since the N-terminal amino acids of Churchill form part of the zinc-binding motif, the addition of a fusion protein at the N terminus causes loss of zinc and unfolding of Churchill. This observation most likely explains the published DNA-binding results, which would arise due to non-specific interaction of the unfolded protein in the immunoprecipitation selection assay. Since Churchill does not appear to bind DNA, we suggest that it may function in embryogenesis as a protein-interaction factor.
- Department of Molecular Biology and Skaggs Institute for Chemical Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: