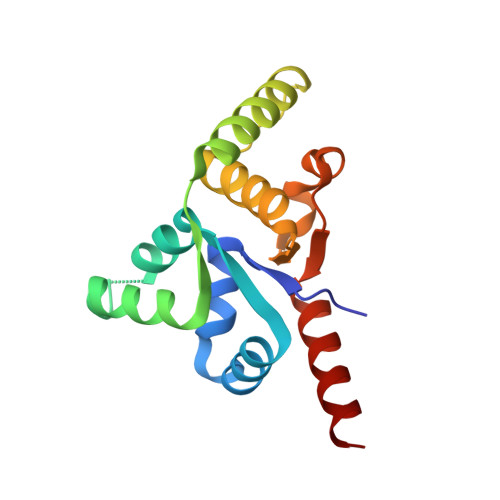
Structure of PIN-domain protein PH0500 from Pyrococcus horikoshii.
Jeyakanthan, J., Inagaki, E., Kuroishi, C., Tahirov, T.H.(2005) Acta Crystallogr Sect F Struct Biol Cryst Commun 61: 463-468
- PubMed: 16511069
- DOI: https://doi.org/10.1107/S1744309105012406
- Primary Citation of Related Structures:
1V96, 1YE5 - PubMed Abstract:
The Pyrococcus horikoshii OT3 protein PH0500 is highly conserved within the Pyrococcus genus of hyperthermophilic archaea and shows low amino-acid sequence similarity with a family of PIN-domain proteins. The protein has been expressed, purified and crystallized in two crystal forms: PH0500-I and PH0500-II. The structure was determined at 2.0 A by the multiple anomalous dispersion method using a selenomethionyl derivative of crystal form PH0500-I (PH0500-I-Se). The structure of PH0500-I has been refined at 1.75 A resolution to an R factor of 20.9% and the structure of PH0500-II has been refined at 2.0 A resolution to an R factor of 23.4%. In both crystal forms as well as in solution the molecule appears to be a dimer. Searches of the databases for protein-fold similarities confirmed that the PH0500 protein is a PIN-domain protein with possible exonuclease activity and involvement in DNA or RNA editing.
- Advanced Protein Crystallography Research Group, RIKEN Harima Institute, 1-1-1 Kouto, Sayo-gun, Hyogo 679-5148, Japan.
Organizational Affiliation: