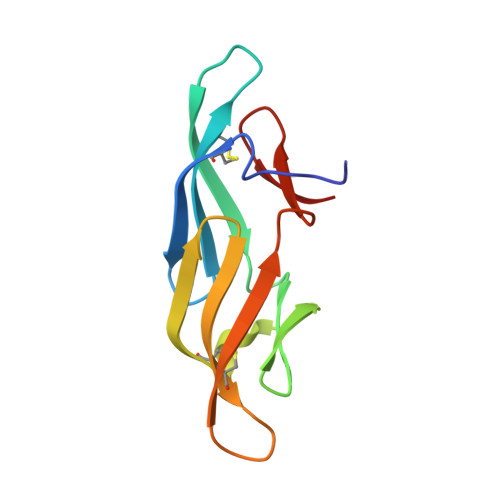
Solution Structure of a Circular-Permuted Variant of the Potent HIV-inactivating Protein Cyanovirin-N: Structural Basis for Protein Stability and Oligosaccharide Interaction
Barrientos, L.G., Louis, J.M., Ratner, D.M., Seeberger, P.H., Gronenborn, A.M.(2003) J Mol Biology 325: 211-223
- PubMed: 12473463
- DOI: https://doi.org/10.1016/s0022-2836(02)01205-6
- Primary Citation of Related Structures:
1N02 - PubMed Abstract:
The high-resolution solution structure of a monomeric circular permuted (cp) variant of the potent HIV-inactivating protein cyanovirin-N (CV-N) was determined by NMR. Comparison with the wild-type (wt) structure revealed that the observed loss in stability of cpCV-N compared to the wt protein is due to less favorable packing of several residues at the pseudo twofold axis that are responsible for holding the two halves of the molecule together. In particular, the N and C-terminal amino acid residues exhibit conformational flexibility, resulting in fewer and less favorable contacts between them. The important hydrophobic and hydrogen-bonding network between residues W49, D89, H90, Y100 and E101 that was observed in wt CV-N is no longer present. For instance, Y100 and E101 are flexible and the tryptophan side-chain is in a different conformation compared to the wt protein. The stability loss amounts to approximately 2kcal/mol and the mobility of the protein is evident by fast amide proton exchange throughout the chain. Mutation of the single proline residue to glycine (P52G) did not substantially affect the stability of the protein, in contrast to the finding for wtCV-N. The binding of high-mannose type oligosaccharides to cpCV-N was also investigated. Similar to wtCV-N, two carbohydrate-binding sites were identified on the protein and the Man alpha1-->2Man linked moieties on the sugar were delineated as binding epitopes. Unlike in wtCV-N, the binding sites on cpCV-N are structurally similar and exhibit comparable binding affinities for the respective sugars. On the basis of the studies presented here and previous results on high-mannose binding to wtCV-N, we discuss a model for the interaction between gp120 and CV-N.
- Laboratory of Chemical Physics, National Institute of Diabetes and Digestive and Kidney Diseases/NIH, Building 5, Room 130, Bethesda, MD 20892-0520, USA.
Organizational Affiliation: