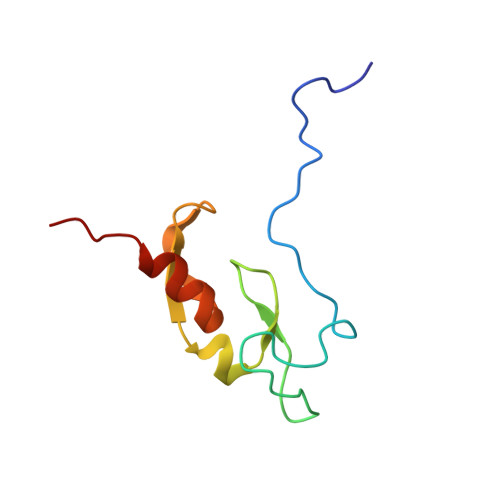
A high-resolution solution structure of a trypanosomatid FYVE domain.
Mertens, H.D.T., Callaghan, J.M., Swarbrick, J.D., McConville, M.J., Gooley, P.R.(2007) Protein Sci 16: 2552-2559
- PubMed: 17905827
- DOI: https://doi.org/10.1110/ps.073009807
- Primary Citation of Related Structures:
1Z2Q - PubMed Abstract:
FYVE domain proteins play key roles in regulating membrane traffic in eukaryotic cells. The FYVE domain displays a remarkable specificity for the head group of the target lipid, phosphatidylinositol 3-phosphate (PtdIns[3]P). We have identified five putative FYVE domain proteins in the genome of the protozoan parasite Leishmania major, three of which are predicted to contain a functional PtdIns(3)P-binding site. The FYVE domain of one of these proteins, LmFYVE-1, bound PtdIns(3)P in liposome-binding assays and targeted GFP to acidified late endosomes/lysosomes in mammalian cells. The high-resolution solution structure of its N-terminal FYVE domain (LmFYVE-1[1-79]) was solved by nuclear magnetic resonance. Functionally significant clusters of residues of the LmFYVE-1 domain involved in PtdIns(3)P binding and dependence on low pH for tight binding were identified. This structure is the first trypanosomatid membrane trafficking protein to be determined and has been refined to high precision and accuracy using residual dipolar couplings.
- Department of Biochemistry and Molecular Biology, Institute of Biotechnology and Molecular Science, University of Melbourne, Parkville, Victoria, Australia.
Organizational Affiliation: