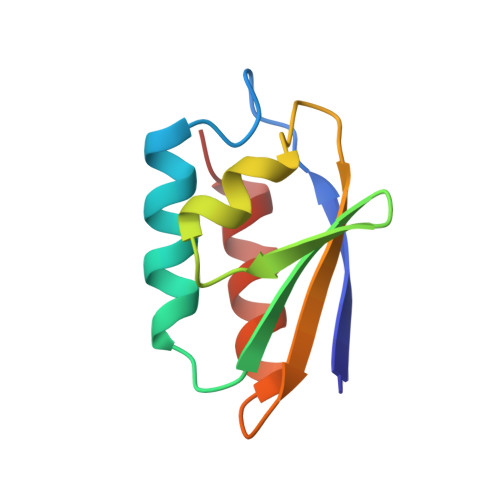
Mutation of serine-46 to aspartate in the histidine-containing protein of Escherichia coli mimics the inactivation by phosphorylation of serine-46 in HPrs from gram-positive bacteria.
Napper, S., Anderson, J.W., Georges, F., Quail, J.W., Delbaere, L.T., Waygood, E.B.(1996) Biochemistry 35: 11260-11267
- PubMed: 8784179
- DOI: https://doi.org/10.1021/bi9603480
- Primary Citation of Related Structures:
1OPD - PubMed Abstract:
Histidine-containing protein (HPr) is a phosphocarrier protein of the bacterial phosphoenolpyruvate:sugar phosphotransferase system. HPr is phosphorylated at the active site residue, His15, by phosphoenolpyruvate-dependent enzyme I in the first enzyme reaction in the process of phosphoryl transfer to sugar. In many Gram-positive bacterial species HPr may also be phosphorylated at Ser46 by an ATP-dependent protein kinase but not in the Gram-negative Escherichia coli and Salmonella typhimurium. One effect of the phosphorylation at Ser46 is to make HPr a poor acceptor for phosphorylation at His15. In Bacillus subtilis HPr, the mutation Ser46Asp mimics the effects of phosphorylation. A series of mutations were made at Ser46 in E. coli HPr: Ala, Arg, Asn, Asp, Glu, and Gly. The two acidic replacements mimic the effects of phosphorylation of Ser46 in HPrs from Gram-positive bacteria. In particular, when mutated to Asp46, the His 15 phosphoacceptor activity (enzyme I Km/Kcat) decreases by about 2000-fold (enzyme I Km, 4 mM HPr; Kcat, approximately 30%). The alanine and glycine mutations had near-wild-type properties, and the asparagine and arginine mutations yielded small changes to the Km values. The crystallographic tertiary structure of Ser46Asp HPr has been determined at 1.5 A resolution, and several changes have been observed which appear to be the effect of the mutation. There is a tightening of helix B, which is demonstrated by a consistent shortening of hydrogen bond lengths throughout the helix as compared to the wild-type structure. There is a repositioning of the Gly54 residue to adopt a 3(10) helical pattern which is not present in the wild-type HPr. In addition, the higher resolution of the mutant structure allows for a more definitive placement of the carbonyl of Pro11. The consequence of this change is that there is no torsion angle strain at residue 16. This result suggests that there is no active site torsion angle strain in wild-type E. coli HPr. The lack of substantial change at the active center of E. coli HPr Ser46Asp HPr suggests that the effect of the Ser46 phosphorylation in HPrs from Gram-positive bacteria is due to an electrostatic interference with enzyme I binding.
- Department of Biochemistry, University of Saskatchewan, Saskatoon, Canada.
Organizational Affiliation: