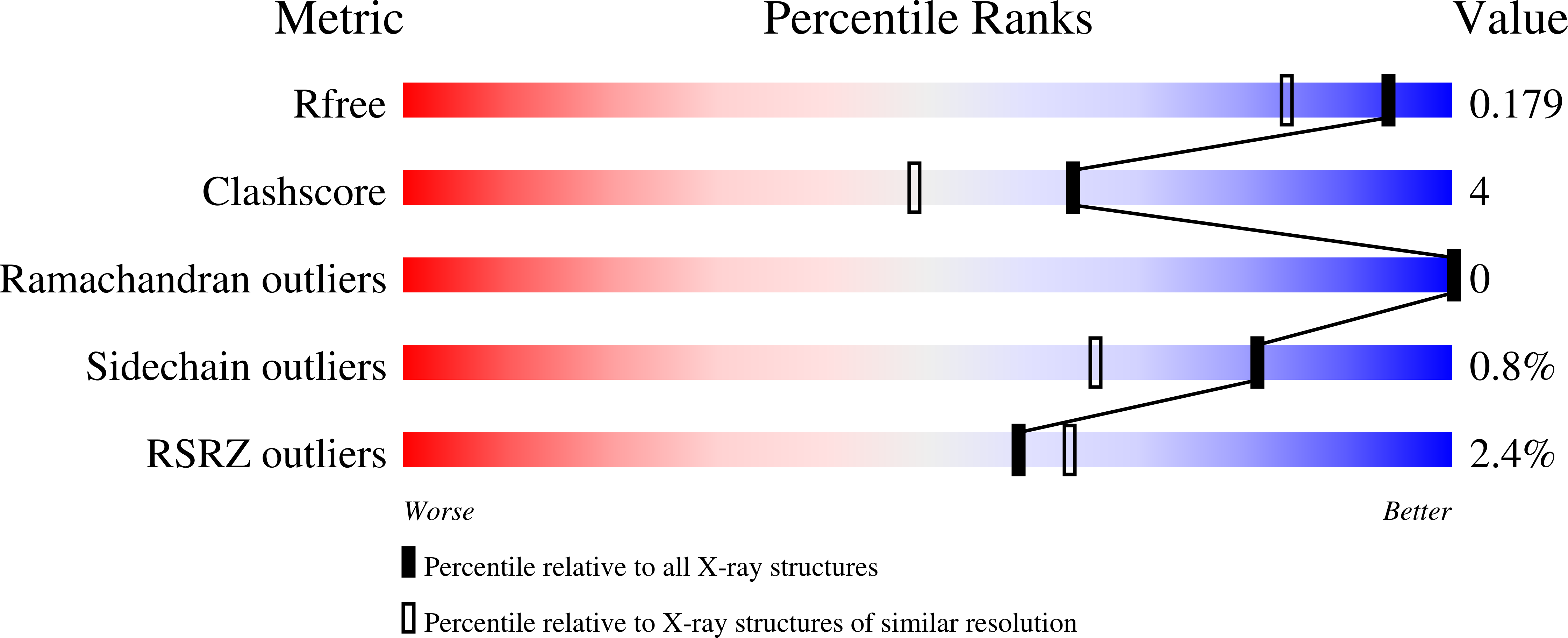
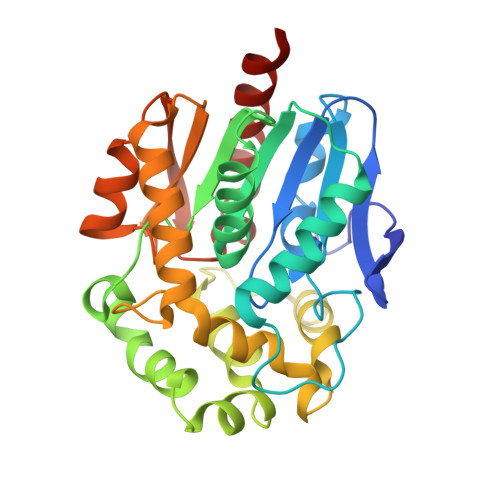
Structural Analysis of the Ancestral Haloalkane Dehalogenase AncLinB-DmbA.
Mazur, A., Grinkevich, P., Chaloupkova, R., Havlickova, P., Kascakova, B., Kuty, M., Damborsky, J., Kuta Smatanova, I., Prudnikova, T.(2021) Int J Mol Sci 22
- PubMed: 34769421
- DOI: https://doi.org/10.3390/ijms222111992
- Primary Citation of Related Structures:
7PW1 - PubMed Abstract:
Haloalkane dehalogenases (EC 3.8.1.5) play an important role in hydrolytic degradation of halogenated compounds, resulting in a halide ion, a proton, and an alcohol. They are used in biocatalysis, bioremediation, and biosensing of environmental pollutants and also for molecular tagging in cell biology. The method of ancestral sequence reconstruction leads to prediction of sequences of ancestral enzymes allowing their experimental characterization. Based on the sequences of modern haloalkane dehalogenases from the subfamily II, the most common ancestor of thoroughly characterized enzymes LinB from Sphingobium japonicum UT26 and DmbA from Mycobacterium bovis 5033/66 was in silico predicted, recombinantly produced and structurally characterized. The ancestral enzyme AncLinB-DmbA was crystallized using the sitting-drop vapor-diffusion method, yielding rod-like crystals that diffracted X-rays to 1.5 Å resolution. Structural comparison of AncLinB-DmbA with their closely related descendants LinB and DmbA revealed some differences in overall structure and tunnel architecture. Newly prepared AncLinB-DmbA has the highest active site cavity volume and the biggest entrance radius on the main tunnel in comparison to descendant enzymes. Ancestral sequence reconstruction is a powerful technique to study molecular evolution and design robust proteins for enzyme technologies.
Organizational Affiliation:
Faculty of Science, University of South Bohemia in Ceske Budejovice, Branisovska 1760, 370 05 Ceske Budejovice, Czech Republic.